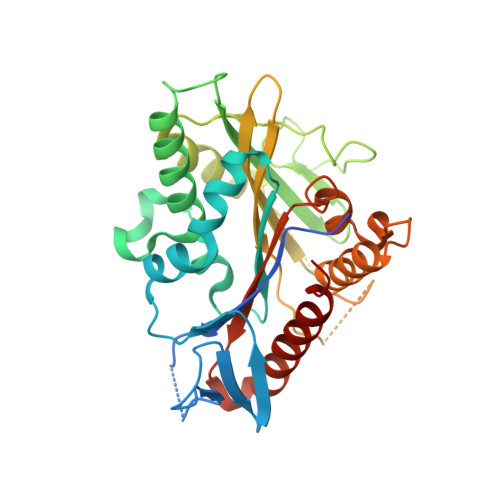
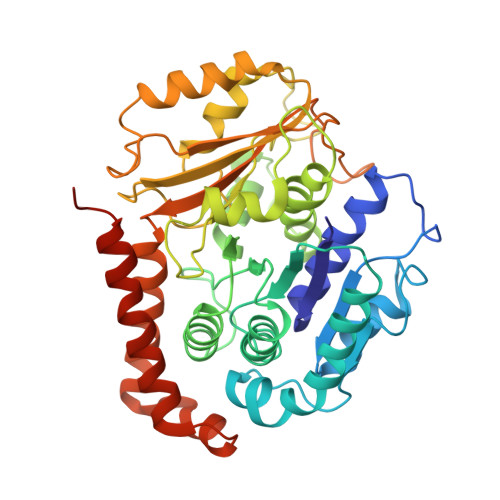
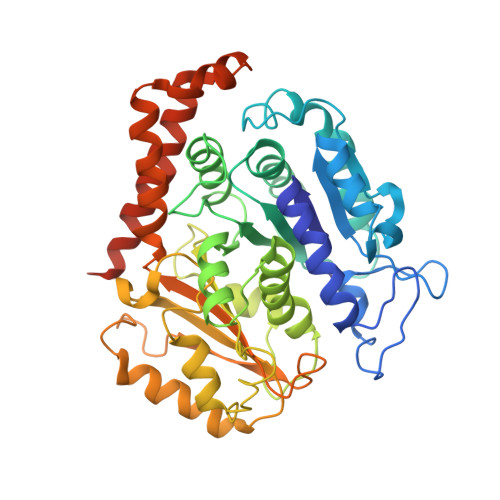
ATPase cycle of the nonmotile kinesin NOD allows microtubule end tracking and drives chromosome movement.
Cochran, J.C., Sindelar, C.V., Mulko, N.K., Collins, K.A., Kong, S.E., Hawley, R.S., Kull, F.J.(2009) Cell 136: 110-122
- PubMed: 19135893 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2008.11.048
- Primary Citation Related Structures:
3DC4, 3DCB, 3DCO - PubMed Abstract:
Segregation of nonexchange chromosomes during Drosophila melanogaster meiosis requires the proper function of NOD, a nonmotile kinesin-10. We have determined the X-ray crystal structure of the NOD catalytic domain in the ADP- and AMPPNP-bound states. These structures reveal an alternate conformation of the microtubule binding region as well as a nucleotide-sensitive relay of hydrogen bonds at the active site. Additionally, a cryo-electron microscopy reconstruction of the nucleotide-free microtubule-NOD complex shows an atypical binding orientation. Thermodynamic studies show that NOD binds tightly to microtubules in the nucleotide-free state, yet other nucleotide states, including AMPPNP, are weakened. Our pre-steady-state kinetic analysis demonstrates that NOD interaction with microtubules occurs slowly with weak activation of ADP product release. Upon rapid substrate binding, NOD detaches from the microtubule prior to the rate-limiting step of ATP hydrolysis, which is also atypical for a kinesin. We propose a model for NOD's microtubule plus-end tracking that drives chromosome movement.
- Department of Chemistry, Dartmouth College, Hanover, NH 03755, USA.
Organizational Affiliation: