Structure and Proposed Activity of a Member of the VapBC Family of Toxin-Antitoxin Systems: VapBC-5 FROM MYCOBACTERIUM TUBERCULOSIS.
Miallau, L., Faller, M., Chiang, J., Arbing, M., Guo, F., Cascio, D., Eisenberg, D.(2009) J Biological Chem 284: 276-283
- PubMed: 18952600 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M805061200
- Primary Citation Related Structures:
3DBO - PubMed Abstract:
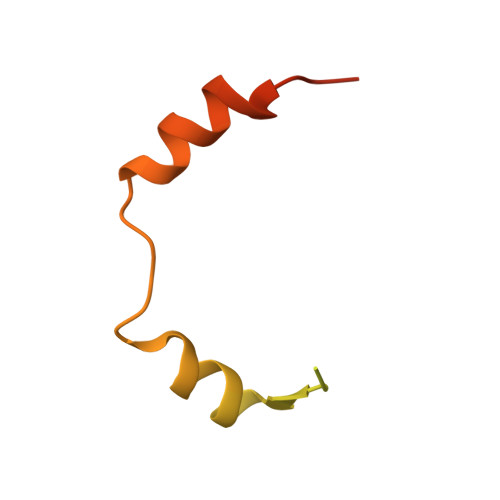
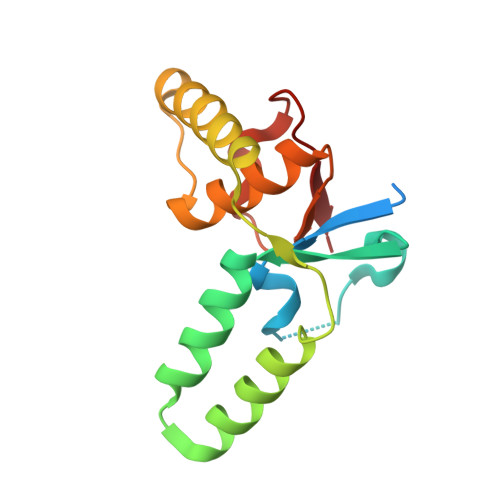
In prokaryotes, cognate toxin-antitoxin pairs have long been known, but no three-dimensional structure has been available for any given complex from Mycobacterium tuberculosis. Here we report the crystal structure and activity of a member of the VapBC family of complexes from M. tuberculosis. The toxin VapC-5 is a compact, 150 residues, two domain alpha/beta protein. Bent around the toxin is the VapB-5 antitoxin, a 33-residue alpha-helix. Assays suggest that the toxin is an Mg-enabled endoribonuclease, inhibited by the antitoxin. The lack of DNase activity is consistent with earlier suggestions that the complex represses its own operon. Furthermore, analysis of the interactions in the binding of the antitoxin to the toxin suggest that exquisite control is required to protect the bacteria cell from toxic VapC-5.
- UCLA-DOE Institute of Genomics and Proteomics, the Department of Biological Chemistry, David Geffen School of Medicine, Molecular Cell and Developmental Biology, and the Department of Chemistry and Biochemistry, University of California, Los Angeles, Los Angeles, California 90095-1570.
Organizational Affiliation: