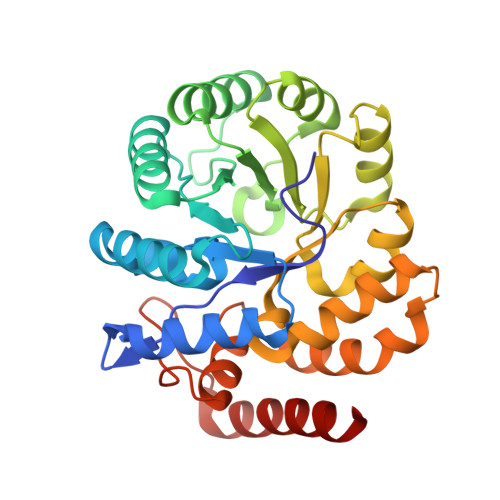
Structure and evolution of a novel dimeric enzyme from a clinically-important bacterial pathogen.
Burgess, B.R., Dobson, R.C.J., Bailey, M.F., Atkinson, S.C., Griffin, M.D.W., Jameson, G.B., Parker, M.W., Gerrard, J.A., Perugini, M.A.(2008) J Biological Chem
- PubMed: 18684709 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M804231200
- Primary Citation Related Structures:
3DAQ - PubMed Abstract:
Dihydrodipicolinate synthase (DHDPS) catalyzes the first committed step of the lysine biosynthetic pathway. The tetrameric structure of DHDPS is thought to be essential for enzymatic activity, as isolated dimeric mutants of Escherichia coli DHDPS possess less than 2.5% that of the activity of the wild-type tetramer. It has recently been proposed that the dimeric form lacks activity due to increased dynamics. Tetramerization, by buttressing two dimers together, reduces dynamics in the dimeric unit and explains why all active bacterial DHDPS enzymes to date have been shown to be homo-tetrameric. However, in this study we demonstrate for the first time that DHDPS from methicillin-resistant Staphylococcus aureus (MRSA) exists in a monomer-dimer equilibrium in solution. Fluorescence-detected analytical ultracentrifugation was employed to show that the dimerization dissociation constant of MRSA-DHDPS is 33 nm in the absence of substrates and 29 nm in the presence of (S)-aspartate semialdehyde (ASA), but is 20-fold tighter in the presence of the substrate pyruvate (1.6 nm). The MRSA-DHDPS dimer exhibits a ping-pong kinetic mechanism (k(cat)=70+/-2 s(-1), K(m)(Pyruvate)=0.11+/-0.01 mm, and K(m)(ASA)=0.22+/-0.02 mm) and shows ASA substrate inhibition with a K(si)(ASA) of 2.7+/-0.9 mm. We also demonstrate that unlike the E. coli tetramer, the MRSA-DHDPS dimer is insensitive to lysine inhibition. The near atomic resolution (1.45 A) crystal structure confirms the dimeric quaternary structure and reveals that the dimerization interface of the MRSA enzyme is more extensive in buried surface area and noncovalent contacts than the equivalent interface in tetrameric DHDPS enzymes from other bacterial species. These data provide a detailed mechanistic insight into DHDPS catalysis and the evolution of quaternary structure of this important bacterial enzyme.
- Bio21 Molecular Science and Biotechnology Institute, University of Melbourne, VIC 3010, Australia; Department of Biochemistry and Molecular Biology, University of Melbourne, VIC 3010, Australia.
Organizational Affiliation: