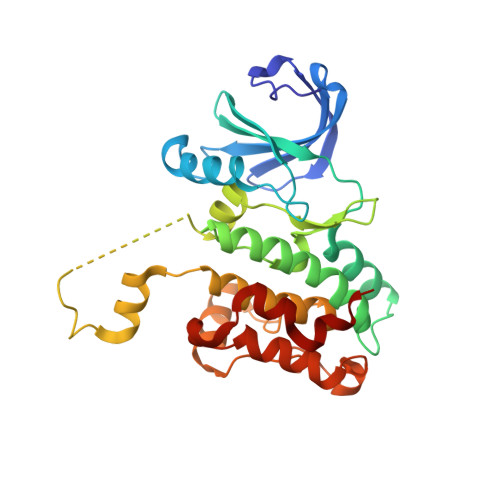
Crystal structure of domain-swapped STE20 OSR1 kinase domain.
Lee, S.J., Cobb, M.H., Goldsmith, E.J.(2008) Protein Sci 18: 304-313
- PubMed: 19177573 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.27
- Primary Citation Related Structures:
3DAK - PubMed Abstract:
OSR1 (oxidative stress-responsive-1) and SPAK (Ste20/Sps1-related proline/alanine-rich kinase) belong to the GCK-VI subfamily of Ste20 group kinases. OSR1 and SPAK are key regulators of NKCCs (Na(+)/K(+)/2Cl(-) cotransporters) and activated by WNK family members (with-no-lysine kinase), mutations of which are known to cause Gordon syndrome, an autosomal dominant form of inherited hypertension. The crystal structure of OSR1 kinase domain has been solved at 2.25 A. OSR1 forms a domain-swapped dimer in an inactive conformation, in which P+1 loop and alphaEF helix are swapped between dimer-related monomers. Structural alignment with nonswapped Ste20 TAO2 kinase indicates that the integrity of chemical interactions in the kinase domain is well preserved in the domain-swapped interfaces. The OSR1 kinase domain has now been added to a growing list of domain-swapped protein kinases recently reported, suggesting that the domain-swapping event provides an additional layer of complexity in regulating protein kinase activity.
- Department of Biochemistry, The University of Texas Southwestern Medical Center at Dallas, 75390-9041, USA.
Organizational Affiliation: