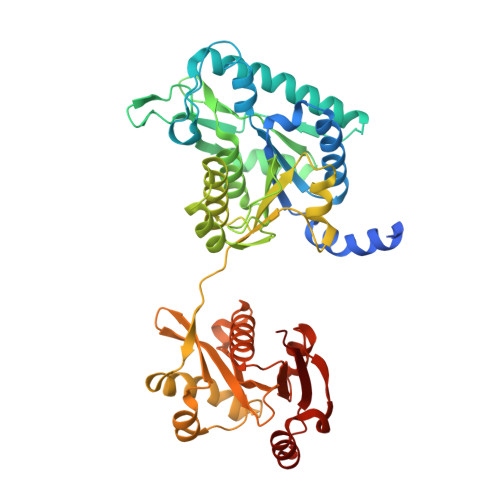
Mechanism of Allosteric Inhibition of N-Acetyl-L-glutamate Synthase by L-Arginine.
Min, L., Jin, Z., Caldovic, L., Morizono, H., Allewell, N.M., Tuchman, M., Shi, D.(2009) J Biological Chem 284: 4873-4880
- PubMed: 19095660 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M805348200
- Primary Citation Related Structures:
3D2M, 3D2P - PubMed Abstract:
N-Acetylglutamate synthase (NAGS) catalyzes the first committed step in l-arginine biosynthesis in plants and micro-organisms and is subject to feedback inhibition by l-arginine. This study compares the crystal structures of NAGS from Neisseria gonorrhoeae (ngNAGS) in the inactive T-state with l-arginine bound and in the active R-state complexed with CoA and l-glutamate. Under all of the conditions examined, the enzyme consists of two stacked trimers. Each monomer has two domains: an amino acid kinase (AAK) domain with an AAK-like fold but lacking kinase activity and an N-acetyltransferase (NAT) domain homologous to other GCN5-related transferases. Binding of l-arginine to the AAK domain induces a global conformational change that increases the diameter of the hexamer by approximately 10 A and decreases its height by approximately 20A(.) AAK dimers move 5A outward along their 2-fold axes, and their tilt relative to the plane of the hexamer decreases by approximately 4 degrees . The NAT domains rotate approximately 109 degrees relative to AAK domains enabling new interdomain interactions. Interactions between AAK and NAT domains on different subunits also change. Local motions of several loops at the l-arginine-binding site enable the protein to close around the bound ligand, whereas several loops at the NAT active site become disordered, markedly reducing enzymatic specific activity.
- Research Center for Genetic Medicine, Children's National Medical Center, The George Washington University, Washington, D. C. 20010, USA.
Organizational Affiliation: