Discovery of benzamide tetrahydro-4H-carbazol-4-ones as novel small molecule inhibitors of Hsp90
Barta, T.E., Veal, J.M., Rice, J.W., Partridge, J.M., Fadden, R.P., Ma, W., Jenks, M., Geng, L., Hanson, G.J., Huang, K.H., Barabasz, A.F., Foley, B.E., Otto, J., Hall, S.E.(2008) Bioorg Med Chem Lett 18: 3517-3521
- PubMed: 18511277 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2008.05.023
- Primary Citation Related Structures:
3D0B - PubMed Abstract:
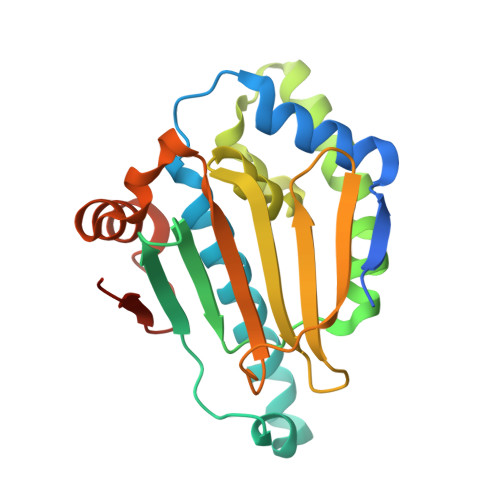
Hsp90 maintains the conformational stability of multiple proteins implicated in oncogenesis and has emerged as a target for chemotherapy. We report here the discovery of a novel small molecule scaffold that inhibits Hsp90. X-ray data show that the scaffold binds competitively at the ATP site on Hsp90. Cellular proliferation and client assays demonstrate that members of the series are able to inhibit Hsp90 at nanomolar concentrations.
- Serenex, Inc., 323 Foster St., Durham, NC 27701, USA.
Organizational Affiliation: