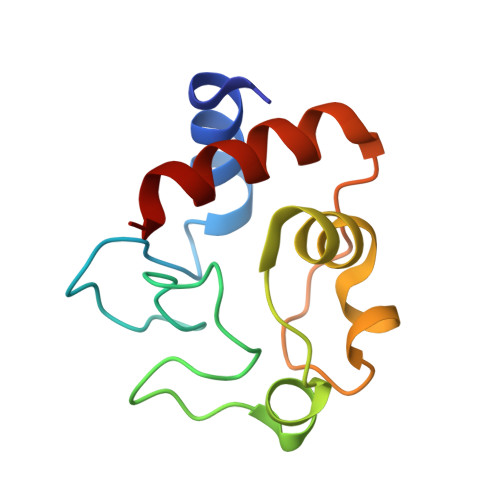
Redox conformation changes in refined tuna cytochrome c.
Takano, T., Dickerson, R.E.(1980) Proc Natl Acad Sci U S A 77: 6371-6375
- PubMed: 6256733 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.77.11.6371
- Primary Citation Related Structures:
3CYT - PubMed Abstract:
Tuna ferrocytochrome c and ferricytochrome c have been refined independently at high resolution (1.5 A and 1.8 A) to crystallographic residual errors of 17.3% and 20.8%, respectively. Small but significant conformational differences are seen surrounding a buried water molecule that is hydrogen bonded to Asn-52, Tyr-67, and Thr-78. In the oxidized state, this water molecule is 1.0 A closer to the heme and the heme has moved 0.15 A out of its heme crevice; both changes lead to a more polar microenvironment for the heme. Chemical modification studies, patterns of evolutionary conservatism, structural differences in bacterial cytochromes, and x-ray studies all agree that the "active site" for cytochrome c is bounded by lysines 8, 13,27, 72, 79, 86, and 87 (thus containing the evolutionary conservative 72-87 loop) and has the buried water molecule just below its surface and the opening of the heme crevice slightly to one side.