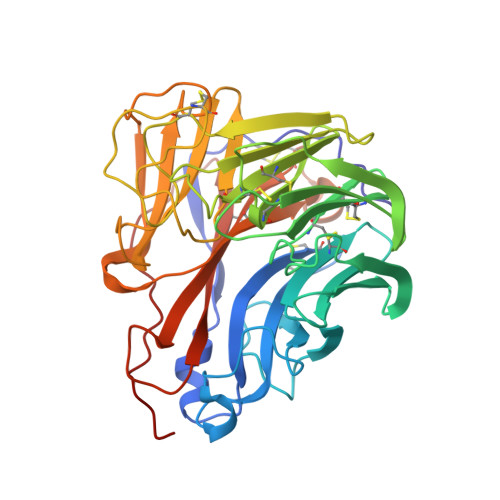
Structure determination of the 1918 H1N1 neuraminidase from a crystal with lattice-translocation defects
Zhu, X., Xu, X., Wilson, I.A.(2008) Acta Crystallogr D Biol Crystallogr 64: 843-850
- PubMed: 18645233 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444908016648
- Primary Citation Related Structures:
3CYE - PubMed Abstract:
Few examples of macromolecular crystals containing lattice-translocation defects have been published in the literature. Lattice translocation and twinning are believed to be two common but different crystal-growth anomalies. While the successful use of twinned data for structure determination has become relatively routine in recent years, structure determination of crystals with lattice-translocation defects has not often been reported. To date, only four protein crystal structures containing such a crystal defect have been determined, using corrected, but not uncorrected, intensity data. In this report, the crystallization, structure determination and refinement of N1 neuraminidase derived from the 1918 H1N1 influenza virus (18NA) at 1.65 A resolution are described. The crystal was indexed in space group C222(1), with unit-cell parameters a = 117.7, b = 138.5, c = 117.9 A, and the structure was solved by molecular replacement. The lattice-translocation vector in the 18NA crystal was (0, 1/2, 1/2) or its equivalent vector (1/2, 0, 1/2) owing to the C lattice symmetry. Owing to this special lattice-translocation vector in space group C222(1), structure refinement could be achieved in two different ways: using corrected or uncorrected diffraction data. In the refinement with uncorrected data, a composite model was built to represent the molecules in the translated and untranslated layers, respectively. This composite structure model provided a unique example to examine how the molecules were arranged in the two lattice domains resulting from lattice-translocation defects.
- Department of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: