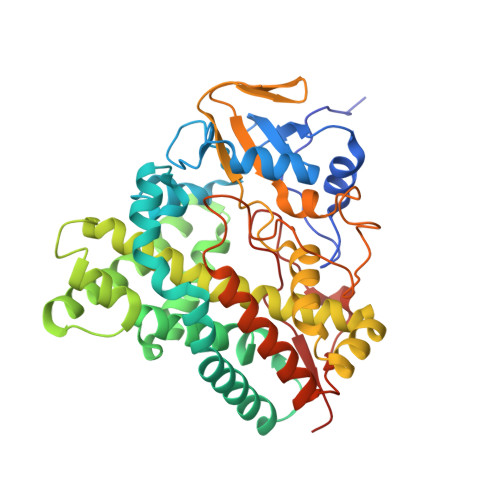
Structure-based design of a highly active vitamin D hydroxylase from Streptomyces griseolus CYP105A1
Hayashi, K., Sugimoto, H., Shinkyo, R., Yamada, M., Ikeda, S., Ikushiro, S., Kamakura, M., Shiro, Y., Sakaki, T.(2008) Biochemistry 47: 11964-11972
- PubMed: 18937506 Search on PubMed
- DOI: https://doi.org/10.1021/bi801222d
- Primary Citation Related Structures:
3CV8, 3CV9 - PubMed Abstract:
CYP105A1 from Streptomyces griseolus has the capability of converting vitamin D 3 (VD 3) to its active form, 1alpha,25-dihydroxyvitamin D 3 (1alpha,25(OH) 2D 3) by a two-step hydroxylation reaction. Our previous structural study has suggested that Arg73 and Arg84 are key residues for the activities of CYP105A1. In this study, we prepared a series of single and double mutants by site-directed mutagenesis focusing on these two residues of CYP105A1 to obtain the hyperactive vitamin D 3 hydroxylase. R84F mutation altered the substrate specificity that gives preference to the 1alpha-hydroxylation of 25-hydroxyvitamin D 3 over the 25-hydroxylation of 1alpha-hydroxyvitamin D 3, opposite to the wild type and other mutants. The double mutant R73V/R84A exhibited 435- and 110-fold higher k cat/ K m values for the 25-hydroxylation of 1alpha-hydroxyvitamin D 3 and 1alpha-hydroxylation of 25-hydroxyvitamin D 3, respectively, compared with the wild-type enzyme. These values notably exceed those of CYP27A1, which is the physiologically essential VD 3 hydroxylase. Thus, we successfully generated useful enzymes of altered substrate preference and hyperactivity. Structural and kinetic analyses of single and double mutants suggest that the amino acid residues at positions 73 and 84 affect the location and conformation of the bound compound in the reaction site and those in the transient binding site, respectively.
- Department of Biotechnology, Faculty of Engineering, Toyama Prefectural University, 5180 Kurokawa, Imizu, Toyama 939-0398, Japan.
Organizational Affiliation: