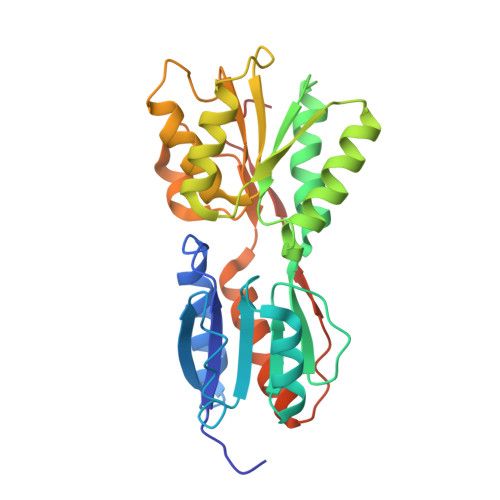
Crystal structure of sugar-binding transcriptional regulator (LacI family) from Enterococcus faecalis.
Patskovsky, Y., Romero, R., Freeman, J., Iizuka, M., Groshong, C., Smith, D., Wasserman, S.R., Sauder, J.M., Burley, S.K., Almo, S.C.To be published.