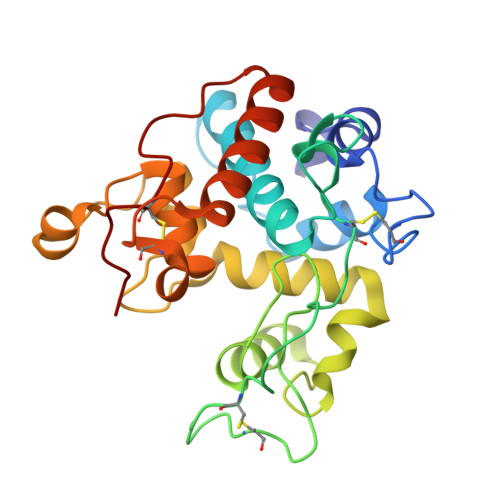
X-ray Structure of Papaya Chitinase Reveals the Substrate Binding Mode of Glycosyl Hydrolase Family 19 Chitinases.
Huet, J., Rucktooa, P., Clantin, B., Azarkan, M., Looze, Y., Villeret, V., Wintjens, R.(2008) Biochemistry 47: 8283-8291
- PubMed: 18636748
- DOI: https://doi.org/10.1021/bi800655u
- Primary Citation of Related Structures:
3CQL - PubMed Abstract:
The crystal structure of a chitinase from Carica papaya has been solved by the molecular replacement method and is reported to a resolution of 1.5 A. This enzyme belongs to family 19 of the glycosyl hydrolases. Crystals have been obtained in the presence of N-acetyl- d-glucosamine (GlcNAc) in the crystallization solution and two well-defined GlcNAc molecules have been identified in the catalytic cleft of the enzyme, at subsites -2 and +1. These GlcNAc moieties bind to the protein via an extensive network of interactions which also involves many hydrogen bonds mediated by water molecules, underlying their role in the catalytic mechanism. A complex of the enzyme with a tetra-GlcNAc molecule has been elaborated, using the experimental interactions observed for the bound GlcNAc saccharides. This model allows to define four major substrate interacting regions in the enzyme, comprising residues located around the catalytic Glu67 (His66 and Thr69), the short segment E89-R90 containing the second catalytic residue Glu89, the region 120-124 (residues Ser120, Trp121, Tyr123, and Asn124), and the alpha-helical segment 198-202 (residues Ile198, Asn199, Gly201, and Leu202). Water molecules from the crystal structure were introduced during the modeling procedure, allowing to pinpoint several additional residues involved in ligand binding that were not previously reported in studies of poly-GlcNAc/family 19 chitinase complexes. This work underlines the role played by water-mediated hydrogen bonding in substrate binding as well as in the catalytic mechanism of the GH family 19 chitinases. Finally, a new sequence motif for family 19 chitinases has been identified between residues Tyr111 and Tyr125.
- Service de Chimie Générale (CP: 206/4), Institut de Pharmacie, Université Libre de Bruxelles (ULB), Campus de la Plaine, Boulevard du Triomphe, B-1050 Brussels, Belgium.
Organizational Affiliation: