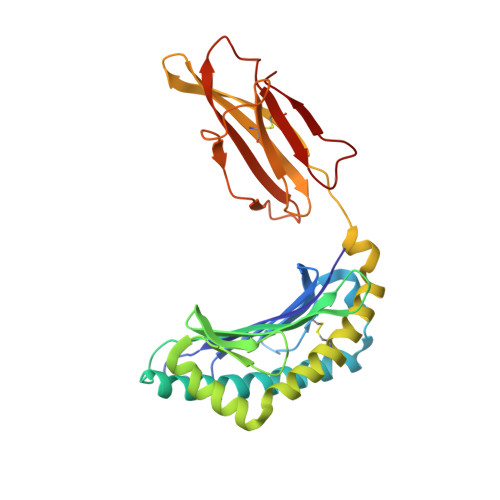
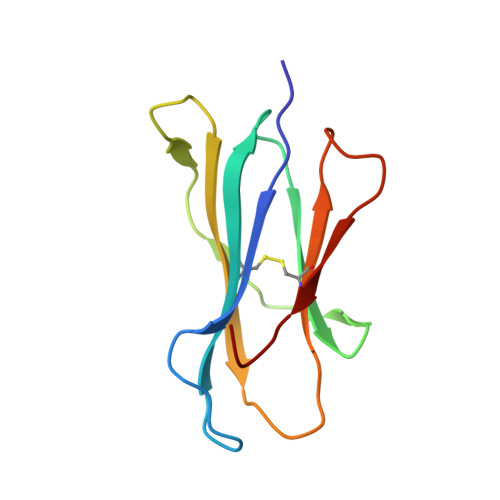
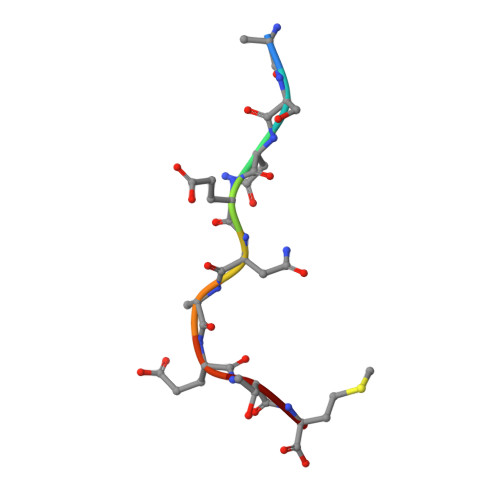
Complete modification of TCR specificity and repertoire selection does not perturb a CD8+ T cell immunodominance hierarchy.
Kedzierska, K., Guillonneau, C., Gras, S., Hatton, L.A., Webby, R., Purcell, A.W., Rossjohn, J., Doherty, P.C., Turner, S.J.(2008) Proc Natl Acad Sci U S A 105: 19408-19413
- PubMed: 19047637 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0810274105
- Primary Citation Related Structures:
3CPL - PubMed Abstract:
Understanding T cell immunodominance hierarchies is fundamental to the development of cellular-based vaccines and immunotherapy. A combination of influenza virus infection in C57BL/6J mice and reverse genetics is used here to dissect the role of T cell antigen receptor (TCR) repertoire in the immunodominant D(b)NP(366)CD8(+) T cell response. Infection with an engineered virus (NPM6A) containing a single alanine (A) mutation at the critical p6 NP(366-374) residue induced a noncross-reactive CD8(+) T cell response characterized by a novel, narrower TCR repertoire per individual mouse that was nonetheless equivalent in magnitude to that generated after WT virus challenge. Although of lower overall avidity, the levels of both cytotoxic T lymphocyte activity and cytokine production were comparable with those seen for the native response. Importantly, the overdominance profile characteristic of secondary D(b)NP(366)-specific clonal expansions was retained for the NPM6A mutant. The primary determinants of immunodominance in this endogenous, non-TCR-transgenic model of viral immunity are thus independent of TCR repertoire composition and diversity. These findings both highlight the importance of effective antigen dose for T cell vaccination and/or immunotherapy and demonstrate the feasibility of priming the memory T cell compartment with engineered viruses to protect against commonly selected mutants viral (or tumor) escape mutants.
- Department of Microbiology and Immunology, The Bio21 Molecular Science and Biotechnology Institute, University of Melbourne, Parkville 3010, Melbourne, Australia. kkedz@unimelb.edu.au
Organizational Affiliation: