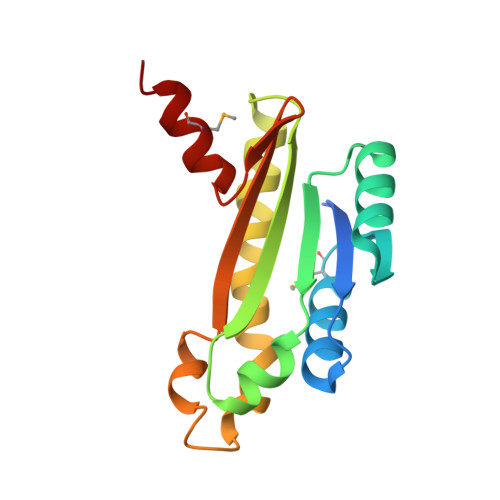
Structural characterization of the putative ABC-type 2 transporter from Thermotoga maritima MSB8.
Filippova, E.V., Tkaczuk, K.L., Chruszcz, M., Xu, X., Savchenko, A., Edwards, A., Minor, W.(2014) J Struct Funct Genomics 15: 215-222
- PubMed: 25306867 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s10969-014-9189-7
- Primary Citation Related Structures:
3CNI - PubMed Abstract:
This study describes the structure of the putative ABC-type 2 transporter TM0543 from Thermotoga maritima MSB8 determined at a resolution of 2.3 Å. In comparative sequence-clustering analysis, TM0543 displays similarity to NatAB-like proteins, which are components of the ABC-type Na(+) efflux pump permease. However, the overall structure fold of the predicted nucleotide-binding domain reveals that it is different from any known structure of ABC-type efflux transporters solved to date. The structure of the putative TM0543 domain also exhibits different dimer architecture and topology of its presumed ATP binding pocket, which may indicate that it does not bind nucleotide at all. Structural analysis of calcium ion binding sites found at the interface between TM0543 dimer subunits suggests that protein may be involved in ion-transporting activity. A detailed analysis of the protein sequence and structure is presented and discussed.
- Department of Molecular Physiology and Biological Physics, University of Virginia, 1340 Jefferson Park Avenue, Charlottesville, VA, 22908, USA.
Organizational Affiliation: