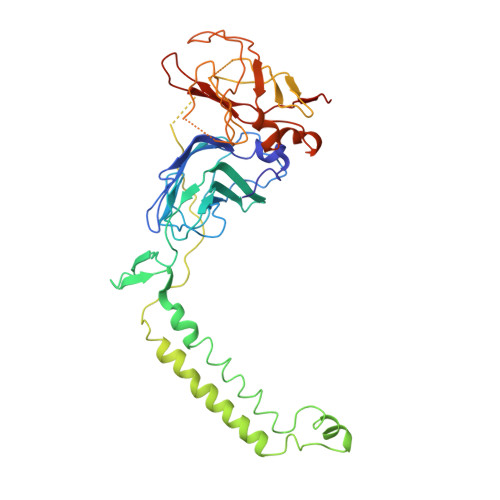
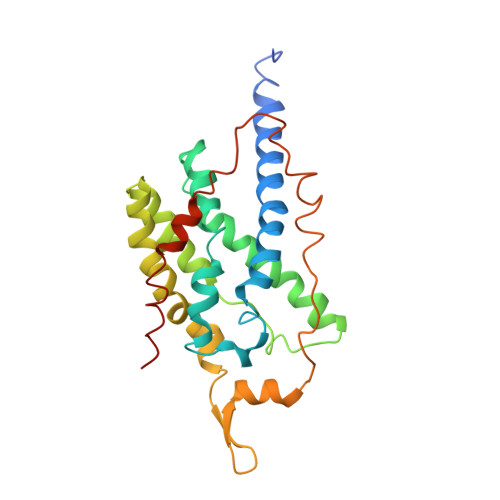
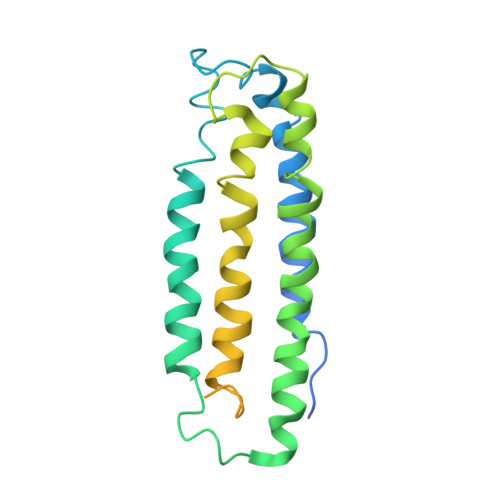
The metal centers of particulate methane monooxygenase from Methylosinus trichosporium OB3b.
Hakemian, A.S., Kondapalli, K.C., Telser, J., Hoffman, B.M., Stemmler, T.L., Rosenzweig, A.C.(2008) Biochemistry 47: 6793-6801
- PubMed: 18540635 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi800598h
- Primary Citation Related Structures:
3CHX - PubMed Abstract:
Particulate methane monooxygenase (pMMO) is a membrane-bound metalloenzyme that oxidizes methane to methanol in methanotrophic bacteria. The nature of the pMMO active site and the overall metal content are controversial, with spectroscopic and crystallographic data suggesting the presence of a mononuclear copper center, a dinuclear copper center, a trinuclear center, and a diiron center or combinations thereof. Most studies have focused on pMMO from Methylococcus capsulatus (Bath). pMMO from a second organism, Methylosinus trichosporium OB3b, has been purified and characterized by spectroscopic and crystallographic methods. Purified M. trichosporium OB3b pMMO contains approximately 2 copper ions per 100 kDa protomer. Electron paramagnetic resonance (EPR) spectroscopic parameters indicate that type 2 Cu(II) is present as two distinct species. Extended X-ray absorption fine structure (EXAFS) data are best fit with oxygen/nitrogen ligands and reveal a Cu-Cu interaction at 2.52 A. Correspondingly, X-ray crystallography of M. trichosporium OB3b pMMO shows a dinuclear copper center, similar to that observed previously in the crystal structure of M. capsulatus (Bath) pMMO. There are, however, significant differences between the pMMO structures from the two organisms. A mononuclear copper center present in M. capsulatus (Bath) pMMO is absent in M. trichosporium OB3b pMMO, whereas a metal center occupied by zinc in the M. capsulatus (Bath) pMMO structure is occupied by copper in M. trichosporium OB3b pMMO. These findings extend previous work on pMMO from M. capsulatus (Bath) and provide new insight into the functional importance of the different metal centers.
- Department of Biochemistry, Northwestern University, Evanston, Illinois 60208, USA.
Organizational Affiliation: