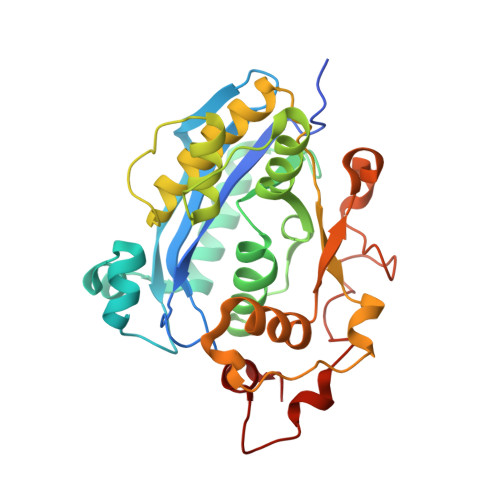
Crystal structure of CRN-4: implications for domain function in apoptotic DNA degradation
Hsiao, Y.-Y., Nakagawa, A., Shi, Z., Mitani, S., Xue, D., Yuan, H.S.(2009) Mol Cell Biol 29: 448-457
- PubMed: 18981218 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/MCB.01006-08
- Primary Citation Related Structures:
3CG7, 3CM5, 3CM6 - PubMed Abstract:
Cell death related nuclease 4 (CRN-4) is one of the apoptotic nucleases involved in DNA degradation in Caenorhabditis elegans. To understand how CRN-4 is involved in apoptotic DNA fragmentation, we analyzed CRN-4's biochemical properties, in vivo cell functions, and the crystal structures of CRN-4 in apo-form, Mn(2+)-bound active form, and Er(3+)-bound inactive form. CRN-4 is a dimeric nuclease with the optimal enzyme activity in cleaving double-stranded DNA in apoptotic salt conditions. Both mutational studies and the structures of the Mn(2+)-bound CRN-4 revealed the geometry of the functional nuclease active site in the N-terminal DEDDh domain. The C-terminal domain, termed the Zn-domain, contains basic surface residues ideal for nucleic acid recognition and is involved in DNA binding, as confirmed by deletion assays. Cell death analysis in C. elegans further demonstrated that both the nuclease active site and the Zn-domain are required for crn-4's function in apoptosis. Combining all of the data, we suggest a structural model where chromosomal DNA is bound at the Zn-domain and cleaved at the DEDDh nuclease domain in CRN-4 when the cell is undergoing apoptosis.
- Institute of Bioinformatics and Structural Biology, National Tsing Hua University, Hsinchu, Taiwan, Republic of China.
Organizational Affiliation: