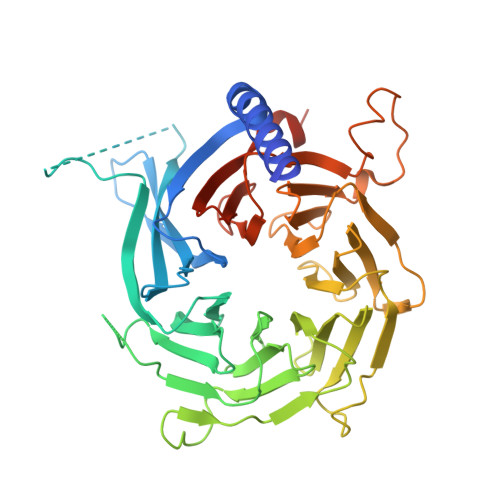
Structural basis of histone H4 recognition by p55.
Song, J.J., Garlick, J.D., Kingston, R.E.(2008) Genes Dev 22: 1313-1318
- PubMed: 18443147
- DOI: https://doi.org/10.1101/gad.1653308
- Primary Citation Related Structures:
3C99, 3C9C - PubMed Abstract:
p55 is a common component of many chromatin-modifying complexes and has been shown to bind to histones. Here, we present a crystal structure of Drosophila p55 bound to a histone H4 peptide. p55, a predicted WD40 repeat protein, recognizes the first helix of histone H4 via a binding pocket located on the side of a beta-propeller structure. The pocket cannot accommodate the histone fold of H4, which must be altered to allow p55 binding. Reconstitution experiments show that the binding pocket is important to the function of p55-containing complexes. These data demonstrate that WD40 repeat proteins use various surfaces to direct the modification of histones.
- Department of Molecular Biology, Massachusetts General Hospital, Boston, Massachusetts 02114, USA.
Organizational Affiliation: