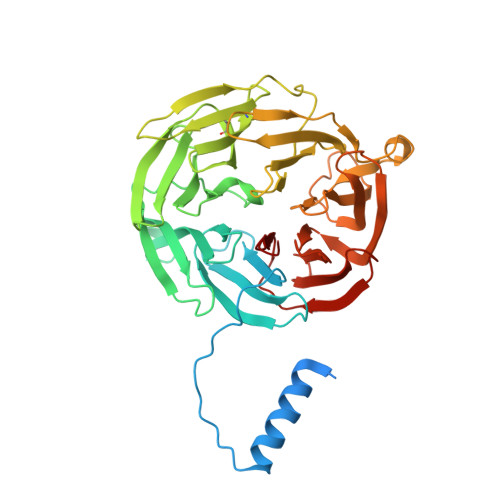
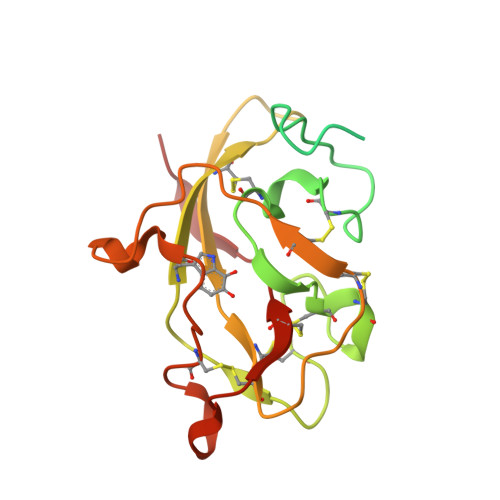
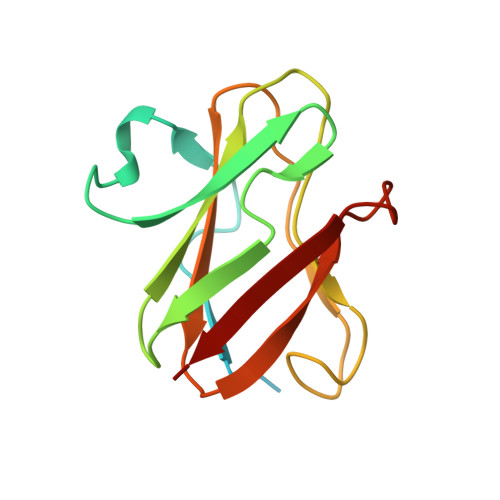
Structural comparison of crystal and solution states of the 138 kDa complex of methylamine dehydrogenase and amicyanin from Paracoccus versutus.
Cavalieri, C., Biermann, N., Vlasie, M.D., Einsle, O., Merli, A., Ferrari, D., Rossi, G.L., Ubbink, M.(2008) Biochemistry 47: 6560-6570
- PubMed: 18512962 Search on PubMed
- DOI: https://doi.org/10.1021/bi7023749
- Primary Citation Related Structures:
3C75 - PubMed Abstract:
Methylamine can be used as the sole carbon source of certain methylotrophic bacteria. Methylamine dehydrogenase catalyzes the conversion of methylamine into formaldehyde and donates electrons to the electron transfer protein amicyanin. The crystal structure of the complex of methylamine dehydrogenase and amicyanin from Paracoccus versutus has been determined, and the rate of electron transfer from the tryptophan tryptophylquinone cofactor of methylamine dehydrogenase to the copper ion of amicyanin in solution has been determined. In the presence of monovalent ions, the rate of electron transfer from the methylamine-reduced TTQ is much higher than in their absence. In general, the kinetics are similar to those observed for the system from Paracoccus denitrificans. The complex in solution has been studied using nuclear magnetic resonance. Signals of perdeuterated, (15)N-enriched amicyanin bound to methylamine dehydrogenase are observed. Chemical shift perturbation analysis indicates that the dissociation rate constant is approximately 250 s(-1) and that amicyanin assumes a well-defined position in the complex in solution. The most affected residues are in the interface observed in the crystal structure, whereas smaller chemical shift changes extend to deep inside the protein. These perturbations can be correlated to small differences in the hydrogen bond network observed in the crystal structures of free and bound amicyanin. This study indicates that chemical shift changes can be used as reliable indicators of subtle structural changes even in a complex larger than 100 kDa.
- Department of Biochemistry and Molecular Biology, University of Parma, 43100 Parma, Italy.
Organizational Affiliation: