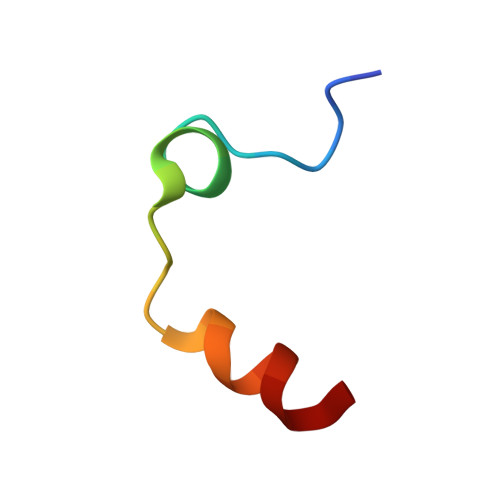
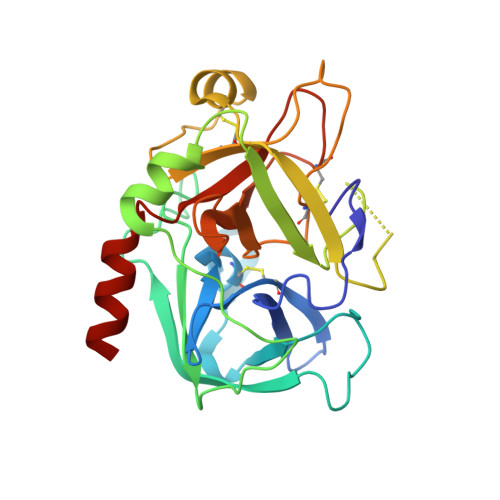
Cyanofluorophenylacetamides as Orally Efficacious Thrombin Inhibitors
Kreuter, K.D., Lee, L., Lu, T., Giardino, E.C., Patel, S., Haung, H., Xu, G., Fitzgerald, M., Mohan, M., Crysler, C., Eisennagel, S., Dasgupta, M., McMillan, M., Spurlino, J.C., Heubert, N., Maryanoff, B.E., Tomczuk, B.E., Damiano, B.P., Player, M.R.To be published.