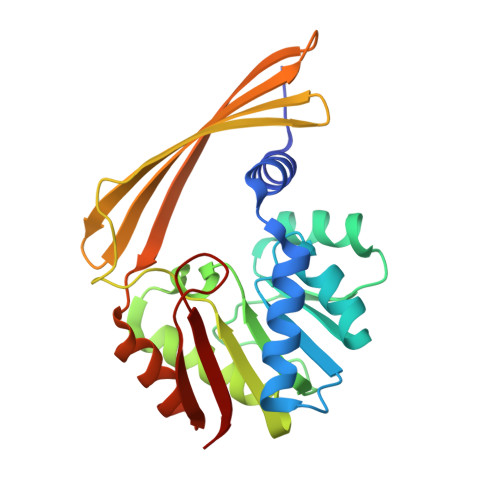
Three-Dimensional Structure of DesVI from Streptomyces venezuelae: A Sugar N,N-Dimethyltransferase Required for dTDP-Desosamine Biosynthesis.
Burgie, E.S., Holden, H.M.(2008) Biochemistry 47: 3982-3988
- PubMed: 18327916
- DOI: https://doi.org/10.1021/bi800063j
- Primary Citation Related Structures:
3BXO - PubMed Abstract:
D-Desosamine, or 3-(dimethylamino)-3,4,6-trideoxyglucose, is an unusual sugar found on the macrolide antibiotic erythromycin, and it has been shown to play a critical role in the biological activity of the drug. Desosamine is added to the parent aglycone via the action of a glycosyltransferase that utilizes dTDP-desosamine as its substrate. Six enzymes are required for the biosynthesis of dTDP-desosamine in Streptomyces venezuelae, with the last step catalyzed by DesVI, an N, N-dimethyltransferase. Here we describe the X-ray crystal structure determined to 2.0 A resolution of DesVI complexed with S-adenosylmethionine (SAM) and the substrate analogue UDP-benzene. Each subunit of the DesVI dimer contains a seven-stranded mixed beta-sheet flanked on either side by alpha-helices. In addition to this major tertiary structural element, there is a four-stranded antiparallel beta-sheet that provides the platform necessary for subunit-subunit assembly. On the basis of the UDP-benzene binding mode, the DesVI substrate, dTDP-3-(methylamino)-3,4,6-trideoxyglucose, has been modeled into the active site. This model places the C-6' methyl group of the sugar into a hydrophobic patch that is well-conserved among putative nucleotide-linked sugar dimethyltransferases. It is formed by Trp 140, Met 178, and Ile 200. The sugar C-2' hydroxyl sits near Tyr 14, and its C-3' amino group is properly positioned for direct in-line attack of the cofactor's reactive methyl group. While methyltransferases that catalyze single alkylations at carbons, oxygens, sulfurs, and nitrogens have been well characterized, little is known regarding enzymes capable of N,N-dimethylation reactions. As such, the ternary structure of DesVI reported here serves as a structural paradigm for a new family of dimethyltransferases that function on nucleotide-linked sugars.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.
Organizational Affiliation: