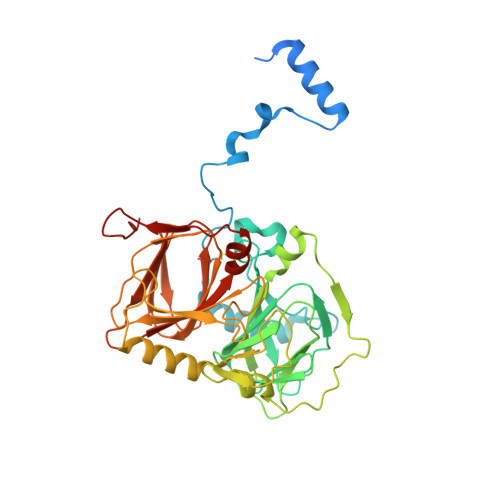
Crystal structure and mutagenic analysis of GDOsp, a gentisate 1,2-dioxygenase from Silicibacter pomeroyi.
Chen, J., Li, W., Wang, M., Zhu, G., Liu, D., Sun, F., Hao, N., Li, X., Rao, Z., Zhang, X.C.(2008) Protein Sci 17: 1362-1373
- PubMed: 18505738 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.035881.108
- Primary Citation Related Structures:
3BU7 - PubMed Abstract:
Dioxygenases catalyze dioxygen incorporation into various organic compounds and play a key role in the complex degradation pathway of mono- and polycyclic aromatic and hetero-aromatic compounds. Here we report the crystal structure of gentisate 1,2-dioxygenase from Silicibacter pomeroyi (GDOsp) at a 2.8 A resolution. The enzyme possessed a conserved three-dimensional structure of the bicupin family, forming a homotetramerization. However, each subunit of GDOsp unusually contained two ferrous centers that were located in its two homologous cupin domains, respectively. Further mutagenic analysis indicated that the enzyme activity of GDOsp depends on the microenvironment in both metal-binding sites. Moreover, homologous structural comparison and functional study on GDOsp variants unveiled a group of functionally essential residues and suggested that the active site of the enzyme is located in the amino-terminal domain, but could be influenced by changes in the carboxyl domain. Therefore, GDOsp may provide a working model for studying long-distance communication within a protein (or its complex).
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, People's Republic of China.
Organizational Affiliation: