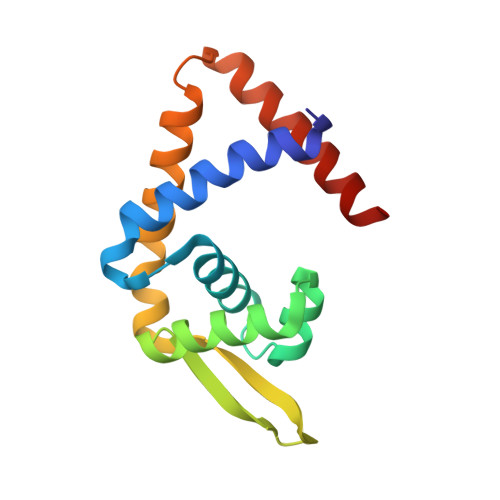
Structural insight on the mechanism of regulation of the MarR family of proteins: high-resolution crystal structure of a transcriptional repressor from Methanobacterium thermoautotrophicum.
Saridakis, V., Shahinas, D., Xu, X., Christendat, D.(2008) J Mol Biology 377: 655-667
- PubMed: 18272181 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.01.001
- Primary Citation Related Structures:
3BPV, 3BPX - PubMed Abstract:
Transcriptional regulators belonging to the MarR family are characterized by a winged-helix DNA binding domain. These transcriptional regulators regulate the efflux and influx of phenolic agents in bacteria and archaea. In Escherichia coli, MarR regulates the multiple antibiotic resistance operon and its inactivation produces a multiple antibiotic resistance phenotype. In some organisms, active efflux of drug compounds will produce a drug resistance phenotype, whereas in other organisms, active influx of chlorinated hydrocarbons results in their rapid degradation. Although proteins in the MarR family are regulators of important biological processes, their mechanism of action is not well understood and structural information about how phenolic agents regulate the activity of these proteins is lacking. This article presents the three-dimensional structure of a protein of the MarR family, MTH313, in its apo form and in complex with salicylate, a known inactivator. A comparison of these two structures indicates that the mechanism of regulation involves a large conformational change in the DNA binding lobe. Electrophoretic mobility shift assay and biophysical analyses further suggest that salicylate inactivates MTH313 and prevents it from binding to its promoter region.
- Department of Biology, York University, 4700 Keele Street, Toronto, Ontario, Canada.
Organizational Affiliation: