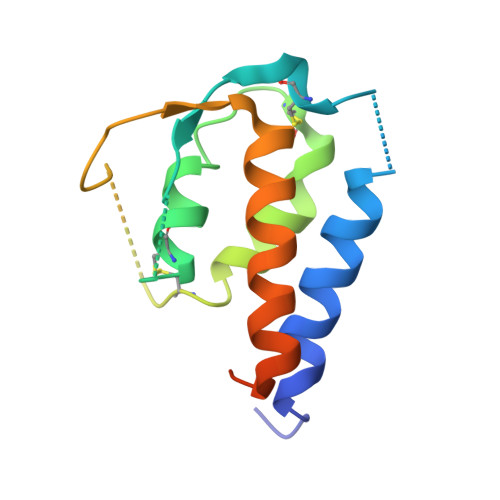
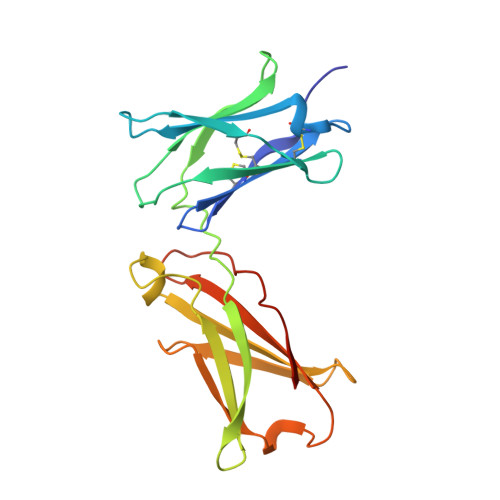
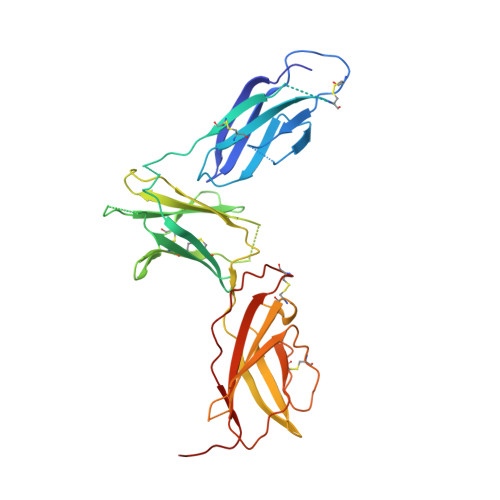
Molecular and Structural Basis of Cytokine Receptor Pleiotropy in the Interleukin-4/13 System.
Laporte, S.L., Juo, Z.S., Vaclavikova, J., Colf, L.A., Qi, X., Heller, N.M., Keegan, A.D., Garcia, K.C.(2008) Cell 132: 259-272
- PubMed: 18243101 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2007.12.030
- Primary Citation Related Structures:
3BPL, 3BPN, 3BPO - PubMed Abstract:
Interleukin-4 and Interleukin-13 are cytokines critical to the development of T cell-mediated humoral immune responses, which are associated with allergy and asthma, and exert their actions through three different combinations of shared receptors. Here we present the crystal structures of the complete set of type I (IL-4R alpha/gamma(c)/IL-4) and type II (IL-4R alpha/IL-13R alpha1/IL-4, IL-4R alpha/IL-13R alpha1/IL-13) ternary signaling complexes. The type I complex reveals a structural basis for gamma(c)'s ability to recognize six different gamma(c)-cytokines. The two type II complexes utilize an unusual top-mounted Ig-like domain on IL-13R alpha1 for a novel mode of cytokine engagement that contributes to a reversal in the IL-4 versus IL-13 ternary complex assembly sequences, which are mediated through substantially different recognition chemistries. We also show that the type II receptor heterodimer signals with different potencies in response to IL-4 versus IL-13 and suggest that the extracellular cytokine-receptor interactions are modulating intracellular membrane-proximal signaling events.
- Howard Hughes Medical Institute, Department of Molecular and Cellular Physiology, Stanford University School of Medicine, Stanford, CA 94305, USA.
Organizational Affiliation: