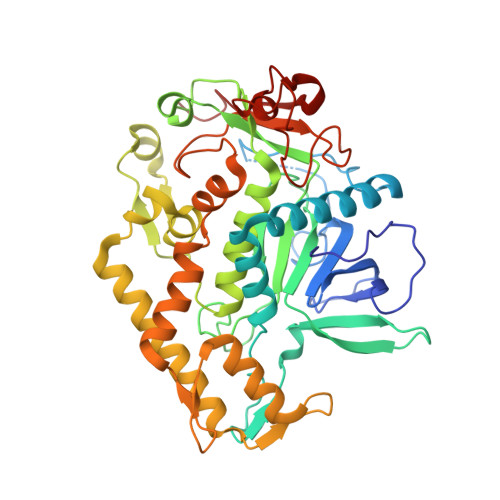
Catalytic features of the botulinum neurotoxin A light chain revealed by high resolution structure of an inhibitory peptide complex.
Silvaggi, N.R., Wilson, D., Tzipori, S., Allen, K.N.(2008) Biochemistry 47: 5736-5745
- PubMed: 18457419
- DOI: https://doi.org/10.1021/bi8001067
- Primary Citation Related Structures:
3BOK, 3BON, 3BOO - PubMed Abstract:
The Clostridium botulinum neurotoxin serotype A light chain (BoNT/A-LC) is a Zn(II)-dependent metalloprotease that blocks the release of acetylcholine at the neuromuscular junction by cleaving SNAP-25, one of the SNARE proteins required for exocytosis. Because of the potential for use of the toxin in bioterrorism and the increasingly widespread application of the toxin in the medical field, there is significant interest in the development of small-molecule inhibitors of the metalloprotease. Efforts to design such inhibitors have not benefited from knowledge of how peptides bind to the active site since the enzyme-peptide structures available previously either were not occupied in the vicinity of the catalytic Zn(II) ion or did not represent the product of SNAP-25 substrate cleavage. Herein we report the 1.4 A-resolution X-ray crystal structure of a complex between the BoNT/A-LC and the inhibitory peptide N-Ac-CRATKML, the first structure of the light chain with an inhibitory peptide bound at the catalytic Zn(II) ion. The peptide is bound with the Cys S gamma atom coordinating the metal ion. Surprisingly, the cysteine sulfur is oxidized to the sulfenic acid form. Given the unstable nature of this species in solution, is it likely that oxidation occurs on the enzyme. In addition to the peptide-bound structure, we report two structures of the unliganded light chain with and without the Zn(II) cofactor bound at 1.25 and 1.20 A resolution, respectively. The two structures are nearly identical, confirming that the Zn(II) ion plays a purely catalytic role. Additionally, the structure of the Zn(II)-bound uncomplexed enzyme allows identification of the catalytic water molecule and a second water molecule that occupies the same position as the peptidic oxygen in the tetrahedral intermediate. This observation suggests that the enzyme active site is prearranged to stabilize the tetrahedral intermediate of the protease reaction.
- Department of Physiology and Biophysics, Boston University School of Medicine, Boston, Massachusetts 02118, USA.
Organizational Affiliation: