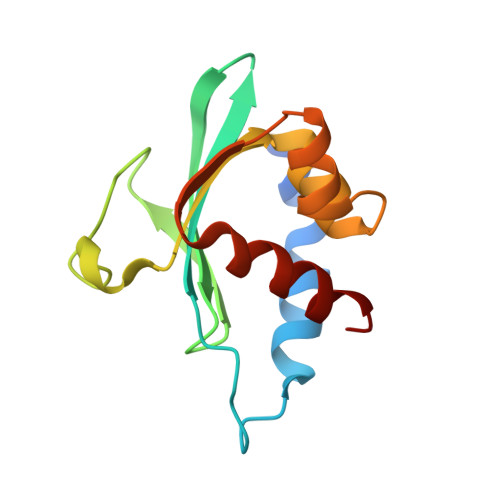
Characterization of the yellow fever mosquito sterol carrier protein-2 like 3 gene and ligand-bound protein structure.
Dyer, D.H., Vyazunova, I., Lorch, J.M., Forest, K.T., Lan, Q.(2009) Mol Cell Biochem 326: 67-77
- PubMed: 19130179 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s11010-008-0007-z
- Primary Citation Related Structures:
3BKR, 3BKS - PubMed Abstract:
The sterol carrier protein-2 like 3 gene (AeSCP-2L3), a new member of the SCP-2 protein family, is identified from the yellow fever mosquito, Aedes aegypti. The predicted molecular weight of AeSCP-2L3 is 13.4 kDa with a calculated pI of 4.98. AeSCP-2L3 transcription occurs in the larval feeding stages and the mRNA levels decrease in pupae and adults. The highest levels of AeSCP-2L3 gene expression are found in the body wall, and possibly originated in the fat body. This is the first report of a mosquito SCP-2-like protein with prominent expression in tissue other than the midgut. The X-ray protein crystal structure of AeSCP-2L3 reveals a bound C16 fatty acid whose acyl tail penetrates deeply into a hydrophobic cavity. Interestingly, the ligand-binding cavity is slightly larger than previously described for AeSCP-2 (Dyer et al. J Biol Chem 278:39085-39091, 2003) and AeSCP-2L2 (Dyer et al. J Lipid Res M700460-JLR200, 2007). There are also an additional 10 amino acids in SCP-2L3 that are not present in other characterized mosquito SCP-2s forming an extended loop between beta 3 and beta 4. Otherwise, the protein backbone is exceedingly similar to other SCP-2 and SCP-2-like proteins. In contrast to this observed high structural homology of members in the mosquito SCP2 family, the amino acid sequence identity between the members is less than 30%. The results from structural analysis imply that there have been evolutionary constraints that favor the SCP-2 C(alpha) backbone fold while the specificity of ligand binding can be altered.
- Department of Bacteriology, University of Wisconsin-Madison, Madison, WI 53706, USA.
Organizational Affiliation: