Synthesis and structure-activity relationships of soluble 8-substituted 4-(2-chlorophenyl)-9-hydroxypyrrolo[3,4-c]carbazole-1,3(2H,6H)-diones as inhibitors of the Wee1 and Chk1 checkpoint kinases.
Smaill, J.B., Lee, H.H., Palmer, B.D., Thompson, A.M., Squire, C.J., Baker, E.N., Booth, R.J., Kraker, A., Hook, K., Denny, W.A.(2008) Bioorg Med Chem Lett 18: 929-933
- PubMed: 18191399
- DOI: https://doi.org/10.1016/j.bmcl.2007.12.046
- Primary Citation of Related Structures:
3BI6, 3BIZ - PubMed Abstract:
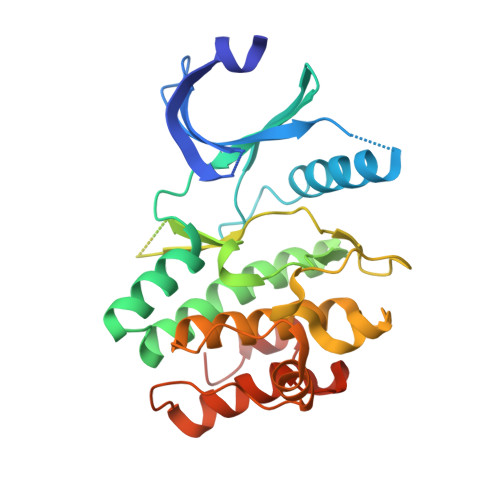
Pyrrolo[3,4-c]carbazoles bearing solubilising basic side chains at the 8-position retain potent Wee1 and Chk1 inhibitory properties in isolated enzyme assays, and evidence of G2/M checkpoint abrogation in several cellular assays. Co-crystal structure studies confirm that the primary binding to the Wee1 enzyme is as described previously, with the C-8 side chains residing in an area of bulk tolerance.
- Auckland Cancer Society Research Centre, School of Medical Sciences, The University of Auckland, Private Bag 92019, Auckland 1142, New Zealand.
Organizational Affiliation: