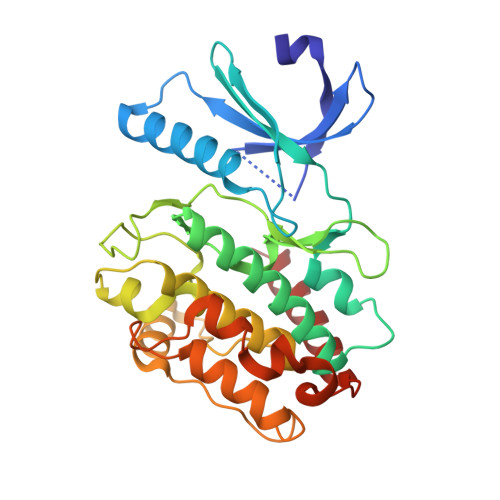
Crystal Structure of Human Calcium/Calmodulin-Dependent Protein Kinase IIB Isoform 1 (CAMK2B).
Filippakopoulos, P., Rellos, P., Niesen, F., Burgess, N., Bullock, A., Berridge, G., Pike, A.C.W., Ugochukwu, E., Pilka, E.S., von Delft, F., Arrowsmith, C.H., Edwards, A.M., Weigelt, J., Knapp, S.To be published.