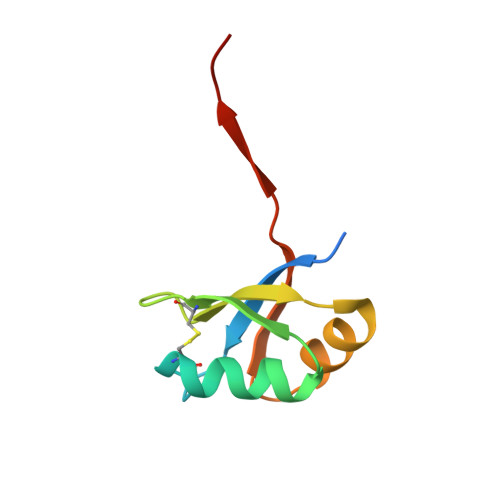
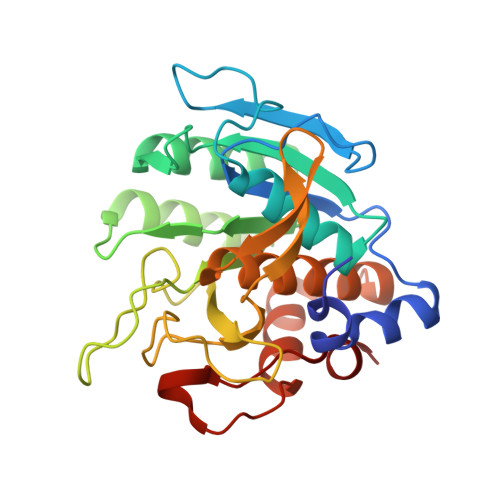
Structure of a switchable subtilisin complexed with a substrate and with the activator azide.
Gallagher, T., Ruan, B., London, M., Bryan, M.A., Bryan, P.N.(2009) Biochemistry 48: 10389-10394
- PubMed: 19761257 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi900577n
- Primary Citation Related Structures:
3BGO, 3CO0 - PubMed Abstract:
An engineered variant of the protease subtilisin from Bacillus amyloliquefaciens, in which the D32A mutation renders the enzyme's activity dependent on the presence of certain small anions such as fluoride or azide, has been produced. This modified enzyme has applications as an azide or fluoride-triggered expression-purification tool. We report activity measurements showing that the enzyme is activated more than 3000-fold by azide and describe the 1.8 A resolution structure of an inactive form (by replacing the catalytic nucleophile Ser 221 with alanine) of the protease, in complex with azide and with a substrate that spans the active site. Both enzyme and substrate have been engineered to increase their stability and the affinity of their interaction. The substrate is based on a stabilized subtilisin prodomain, extended across the active site by the addition of four residues at its C-terminus. In the crystal structure, the substrate is well-ordered across the active site, and the azide anion is observed bound adjacent to Ala 32. The structures of the substrate complex in three different crystals (anion-free, fluoride-soaked, and azide-soaked) are compared. These structures provide extensive information for understanding subtilisin's substrate binding and catalytic mechanism, and for the development of biotechnology tools based on anion-activated proteolysis. The mechanism of anion-dependent proteolysis appears to be a slight modification of the accepted charge-relay mechanism for serine proteases.
- Center for Advanced Research in Biotechnology, 9600 Gudelsky Drive, Rockville, Maryland 20850, USA. gallaghe@umbi.umd.edu
Organizational Affiliation: