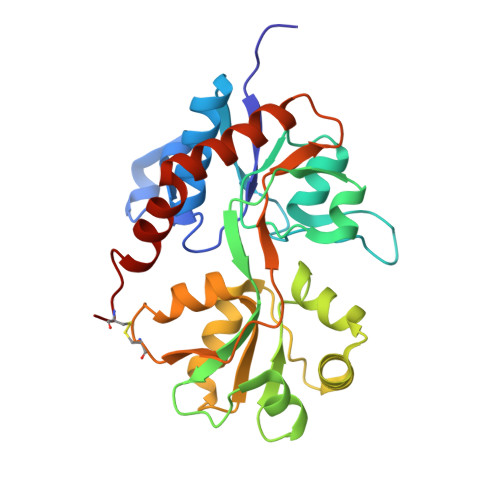
Structures of the ligand-binding core of iGluR2 in complex with the agonists (R)- and (S)-2-amino-3-(4-hydroxy-1,2,5-thiadiazol-3-yl)propionic acid explain their unusual equipotency.
Beich-Frandsen, M., Pickering, D.S., Mirza, O., Johansen, T.N., Greenwood, J., Vestergaard, B., Schousboe, A., Gajhede, M., Liljefors, T., Kastrup, J.S.(2008) J Med Chem 51: 1459-1463
- PubMed: 18269227 Search on PubMed
- DOI: https://doi.org/10.1021/jm701126w
- Primary Citation Related Structures:
3BFT, 3BFU - PubMed Abstract:
AMPA-type ionotropic glutamate receptors generally display high stereoselectivity in agonist binding. However, the stereoisomers of 2-amino-3-(4-hydroxy-1,2,5-thiadiazol-3-yl)propionic acid (TDPA) have similar enantiopharmacology. To understand this observation, we have determined the X-ray structures of ( R)-TDPA and ( S)-TDPA in complex with the ligand-binding core of iGluR2 and investigated the binding pharmacology at AMPA and kainate receptors. Both enantiomers induce full domain closure in iGluR2 but adopt different conformations when binding to the receptor, which may explain the similar enantiopharmacology.
- Department of Medicinal Chemistry, University of Copenhagen, Copenhagen, Denmark.
Organizational Affiliation: