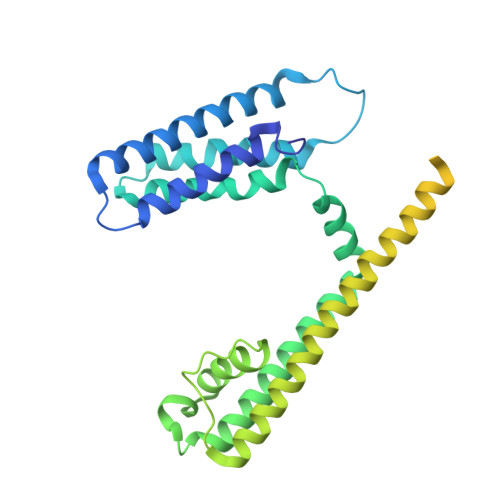
Structure of the transmembrane regions of a bacterial cyclic nucleotide-regulated channel.
Clayton, G.M., Altieri, S., Heginbotham, L., Unger, V.M., Morais-Cabral, J.H.(2008) Proc Natl Acad Sci U S A 105: 1511-1515
- PubMed: 18216238 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0711533105
- Primary Citation Related Structures:
2ZD9, 3BEH - PubMed Abstract:
The six-transmembrane helix (6 TM) tetrameric cation channels form the largest ion channel family, some members of which are voltage-gated and others are not. There are no reported channel structures to match the wealth of functional data on the non-voltage-gated members. We determined the structure of the transmembrane regions of the bacterial cyclic nucleotide-regulated channel MlotiK1, a non-voltage-gated 6 TM channel. The structure showed how the S1-S4 domain and its associated linker can serve as a clamp to constrain the gate of the pore and possibly function in concert with ligand-binding domains to regulate the opening of the pore. The structure also led us to hypothesize a new mechanism by which motions of the S6 inner helices can gate the ion conduction pathway at a position along the pore closer to the selectivity filter than the canonical helix bundle crossing.
- Department of Molecular Biophysics and Biochemistry, Yale University, 260 Whitney Avenue, New Haven, CT 06520, USA.
Organizational Affiliation: