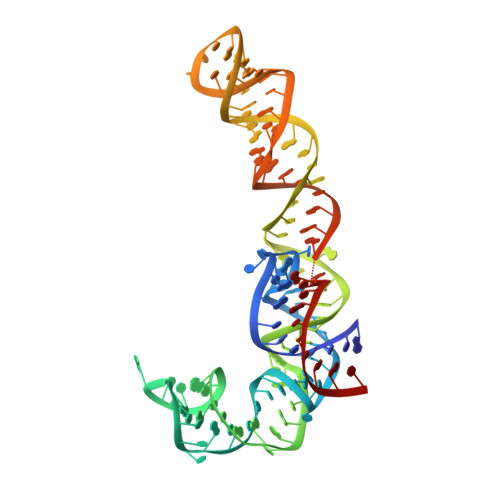
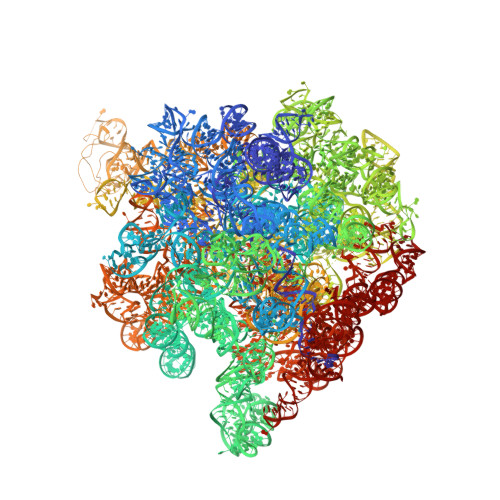
Recycling of Aborted Ribosomal 50S Subunit-Nascent Chain-tRNA Complexes by the Heat Shock Protein Hsp15.
Jiang, L., Schaffitzel, C., Bingel-Erlenmeyer, R., Ban, N., Korber, P., Koning, R.I., de Geus, D.C., Plaisier, J.R., Abrahams, J.P.(2009) J Mol Biology 386: 1357-1367
- PubMed: 19013177 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.10.079
- Primary Citation Related Structures:
3BBU, 3BBV, 3BBX - PubMed Abstract:
When heat shock prematurely dissociates a translating bacterial ribosome, its 50S subunit is prevented from reinitiating protein synthesis by tRNA covalently linked to the unfinished protein chain that remains threaded through the exit tunnel. Hsp15, a highly upregulated bacterial heat shock protein, reactivates such dead-end complexes. Here, we show with cryo-electron microscopy reconstructions and functional assays that Hsp15 translocates the tRNA moiety from the A site to the P site of stalled 50S subunits. By stabilizing the tRNA in the P site, Hsp15 indirectly frees up the A site, allowing a release factor to land there and cleave off the tRNA. Such a release factor must be stop codon independent, suggesting a possible role for a poorly characterized class of putative release factors that are upregulated by cellular stress, lack a codon recognition domain and are conserved in eukaryotes.
- Department of Biophysical Structural Chemistry, Leiden Institute of Chemistry, Leiden University, Einsteinweg 55, 2333 CC Leiden, The Netherlands.
Organizational Affiliation: