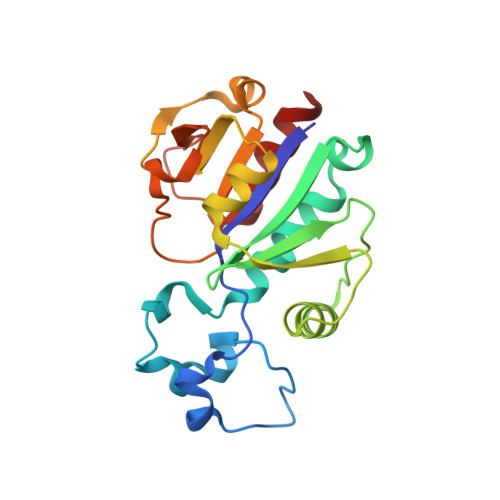
The crystal structure of Nep1 reveals an extended SPOUT-class methyltransferase fold and a pre-organized SAM-binding site.
Taylor, A.B., Meyer, B., Leal, B.Z., Kotter, P., Schirf, V., Demeler, B., Hart, P.J., Entian, K.D., Wohnert, J.(2008) Nucleic Acids Res 36: 1542-1554
- PubMed: 18208838
- DOI: https://doi.org/10.1093/nar/gkm1172
- Primary Citation of Related Structures:
3BBD, 3BBE, 3BBH - PubMed Abstract:
Ribosome biogenesis in eukaryotes requires the participation of a large number of ribosome assembly factors. The highly conserved eukaryotic nucleolar protein Nep1 has an essential but unknown function in 18S rRNA processing and ribosome biogenesis. In Saccharomyces cerevisiae the malfunction of a temperature-sensitive Nep1 protein (nep1-1(ts)) was suppressed by the addition of S-adenosylmethionine (SAM). This suggests the participation of Nep1 in a methyltransferase reaction during ribosome biogenesis. In addition, yeast Nep1 binds to a 6-nt RNA-binding motif also found in 18S rRNA and facilitates the incorporation of ribosomal protein Rps19 during the formation of pre-ribosomes. Here, we present the X-ray structure of the Nep1 homolog from the archaebacterium Methanocaldococcus jannaschii in its free form (2.2 A resolution) and bound to the S-adenosylmethionine analog S-adenosylhomocysteine (SAH, 2.15 A resolution) and the antibiotic and general methyltransferase inhibitor sinefungin (2.25 A resolution). The structure reveals a fold which is very similar to the conserved core fold of the SPOUT-class methyltransferases but contains a novel extension of this common core fold. SAH and sinefungin bind to Nep1 at a preformed binding site that is topologically equivalent to the cofactor-binding site in other SPOUT-class methyltransferases. Therefore, our structures together with previous genetic data suggest that Nep1 is a genuine rRNA methyltransferase.
- Department of Biochemistry, X-ray Crystallography Core Laboratory, The University of Texas Health Science Center San Antonio, San Antonio, TX-78229, USA.
Organizational Affiliation: