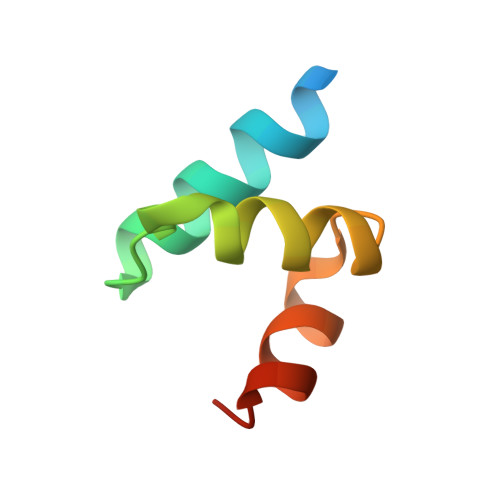
Crystal structure of the ubiquitin-associated (UBA) domain of p62 and its interaction with ubiquitin.
Isogai, S., Morimoto, D., Arita, K., Unzai, S., Tenno, T., Hasegawa, J., Sou, Y.S., Komatsu, M., Tanaka, K., Shirakawa, M., Tochio, H.(2011) J Biological Chem 286: 31864-31874
- PubMed: 21715324
- DOI: https://doi.org/10.1074/jbc.M111.259630
- Primary Citation of Related Structures:
3B0F - PubMed Abstract:
p62/SQSTM1/A170 is a multimodular protein that is found in ubiquitin-positive inclusions associated with neurodegenerative diseases. Recent findings indicate that p62 mediates the interaction between ubiquitinated proteins and autophagosomes, leading these proteins to be degraded via the autophagy-lysosomal pathway. This ubiquitin-mediated selective autophagy is thought to begin with recognition of the ubiquitinated proteins by the C-terminal ubiquitin-associated (UBA) domain of p62. We present here the crystal structure of the UBA domain of mouse p62 and the solution structure of its ubiquitin-bound form. The p62 UBA domain adopts a novel dimeric structure in crystals, which is distinctive from those of other UBA domains. NMR analyses reveal that in solution the domain exists in equilibrium between the dimer and monomer forms, and binding ubiquitin shifts the equilibrium toward the monomer to form a 1:1 complex between the UBA domain and ubiquitin. The dimer-to-monomer transition is associated with a structural change of the very C-terminal end of the p62 UBA domain, although the UBA fold itself is essentially maintained. Our data illustrate that dimerization and ubiquitin binding of the p62 UBA domain are incompatible with each other. These observations reveal an autoinhibitory mechanism in the p62 UBA domain and suggest that autoinhibition plays a role in the function of p62.
- Department of Molecular Engineering, Graduate School of Engineering, Kyoto University, Kyoto-Daigaku Katsura, Kyoto 615-8510, Japan.
Organizational Affiliation: