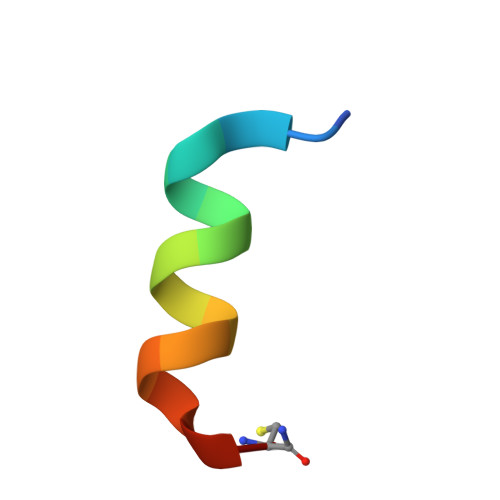
Crystallographic snapshots of tom20-mitochondrial presequence interactions with disulfide-stabilized peptides.
Saitoh, T., Igura, M., Miyazaki, Y., Ose, T., Maita, N., Kohda, D.(2011) Biochemistry 50: 5487-5496
- PubMed: 21591667
- DOI: https://doi.org/10.1021/bi200470x
- Primary Citation of Related Structures:
3AWR, 3AX2, 3AX3, 3AX5 - PubMed Abstract:
Most mitochondrial proteins are synthesized in the cytosol and imported into mitochondria. The Tom20 protein, residing on the mitochondrial surface, recognizes the N-terminal presequences of precursor proteins. We previously determined the crystal structures of the Tom20-presequence complex. The successful crystallization involved tethering the presequence to Tom20 through an intermolecular disulfide bond with an optimized linker. In this work, we assessed the tethering method. The intermolecular disulfide bond was cleaved in crystal with a reducing agent. The pose (i.e., conformation and position) of the presequence was identical to the previously determined pose. In another experiment, a longer linker than the optimized length was used for the tethering. The perturbation of the tether changed the pose slightly, but the interaction mode was preserved. These results argue against the forced interaction of the presequence by its covalent attachment to Tom20. Second, as an alternative method referred to as "molecular stiffening", we introduced a disulfide bond within the presequence peptide to restrict the freedom of the peptide in the unbound states. One presequence analogue exhibited over 100-fold higher affinity than its linear counterpart and generated cocrystals with Tom20. One of the two crystallographic snapshots revealed a known pose previously determined by the tethering method, and the other snapshot depicted a new pose. These results confirmed and extended the dynamic, multiple bound state model of the Tom20-presequence interactions and also demonstrated the validity of the molecular tethering and stiffening techniques in studies of transient protein-peptide interactions.
- Division of Structural Biology, Medical Institute of Bioregulation, Kyushu University, Maidashi 3-1-1, Higashi-ku, Fukuoka 812-8582, Japan.
Organizational Affiliation: