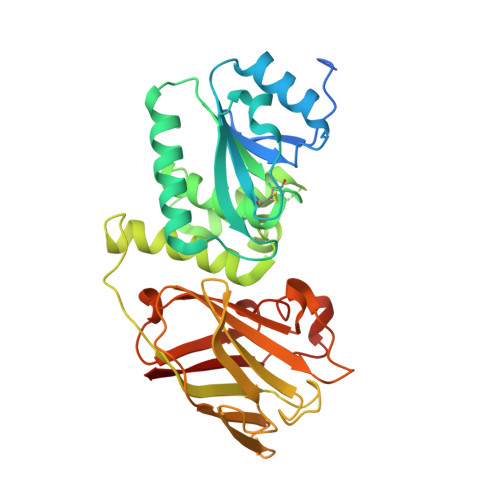
Crystal structure of the cytoplasmic phosphatase and tensin homolog (PTEN)-like region of Ciona intestinalis voltage-sensing phosphatase provides insight into substrate specificity and redox regulation of the phosphoinositide phosphatase activity
Matsuda, M., Takeshita, K., Kurokawa, T., Sakata, S., Suzuki, M., Yamashita, E., Okamura, Y., Nakagawa, A.(2011) J Biological Chem 286: 23368-23377
- PubMed: 21543329 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.214361
- Primary Citation Related Structures:
3AWE, 3AWF, 3AWG - PubMed Abstract:
Ciona intestinalis voltage-sensing phosphatase (Ci-VSP) has a transmembrane voltage sensor domain and a cytoplasmic region sharing similarity to the phosphatase and tensin homolog (PTEN). It dephosphorylates phosphatidylinositol 4,5-bisphosphate and phosphatidylinositol 3,4,5-trisphosphate upon membrane depolarization. The cytoplasmic region is composed of a phosphatase domain and a putative membrane interaction domain, C2. Here we determined the crystal structures of the Ci-VSP cytoplasmic region in three distinct constructs, wild-type (248-576), wild-type (236-576), and G365A mutant (248-576). The crystal structure of WT-236 and G365A-248 had the disulfide bond between the catalytic residue Cys-363 and the adjacent residue Cys-310. On the other hand, the disulfide bond was not present in the crystal structure of WT-248. These suggest the possibility that Ci-VSP is regulated by reactive oxygen species as found in PTEN. These structures also revealed that the conformation of the TI loop in the active site of the Ci-VSP cytoplasmic region was distinct from the corresponding region of PTEN; Ci-VSP has glutamic acid (Glu-411) in the TI loop, orienting toward the center of active site pocket. Mutation of Glu-411 led to acquirement of increased activity toward phosphatidylinositol 3,5-bisphosphate, suggesting that this site is required for determining substrate specificity. Our results provide the basic information of the enzymatic mechanism of Ci-VSP.
- Institute for Protein Research, Graduate School of Medicine, Osaka University, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: