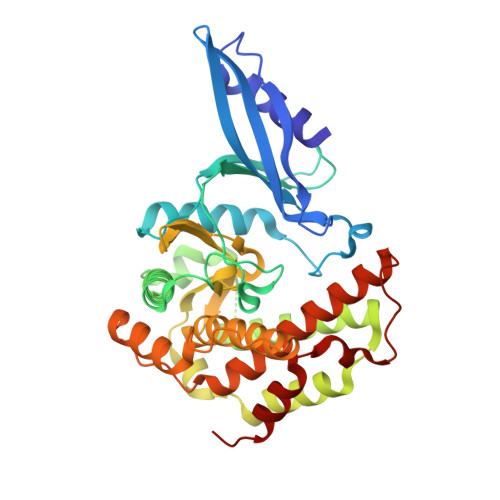
Crystal structure of Mycobacterium tuberculosis Rv3168: a putative aminoglycoside antibiotics resistance enzyme
Kim, S., Nguyen, C.M.T., Kim, E.-J., Kim, K.-J.(2011) Proteins 79: 2983-2987
- PubMed: 21905120 Search on PubMed
- DOI: https://doi.org/10.1002/prot.23119
- Primary Citation Related Structures:
3ATS, 3ATT - Pohang Accelerator Laboratory, Pohang University of Science and Technology, Pohang, Kyungbuk, Korea.
Organizational Affiliation: