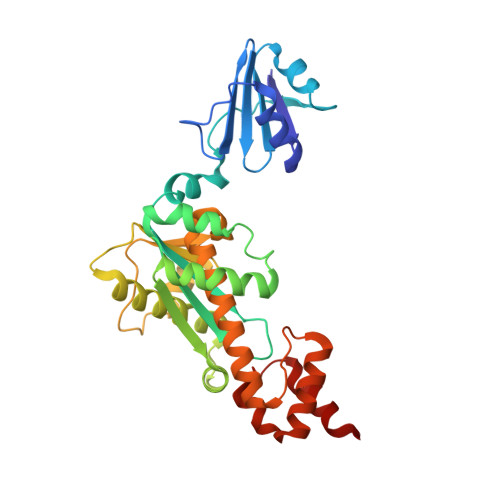
Identification of the substrate binding site in the N-terminal TBP-like domain of RNase H3.
Miyashita, S., Tadokoro, T., Angkawidjaja, C., You, D.-J., Koga, Y., Takano, K., Kanaya, S.(2011) FEBS Lett 585: 2313-2317
- PubMed: 21664908 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2011.05.064
- Primary Citation Related Structures:
3ASM - PubMed Abstract:
Ribonuclease H3 from Bacillus stearothermophilus (Bst-RNase H3) has the N-terminal TBP-like substrate-binding domain. To identify the substrate binding site in this domain, the mutant proteins of the intact protein and isolated N-domain, in which six of the seventeen residues corresponding to those involved in DNA binding of TBP are individually mutated to Ala, were constructed. All of them exhibited decreased enzymatic activities and/or substrate-binding affinities when compared to those of the parent proteins, suggesting that the N-terminal domain of RNase H3 uses the flat surface of the β-sheet for substrate binding as TBP to bind DNA. This domain may greatly change conformation upon substrate binding.
- Department of Material and Life Science, Graduate School of Engineering, Osaka University, Suita, Osaka, Japan.
Organizational Affiliation: