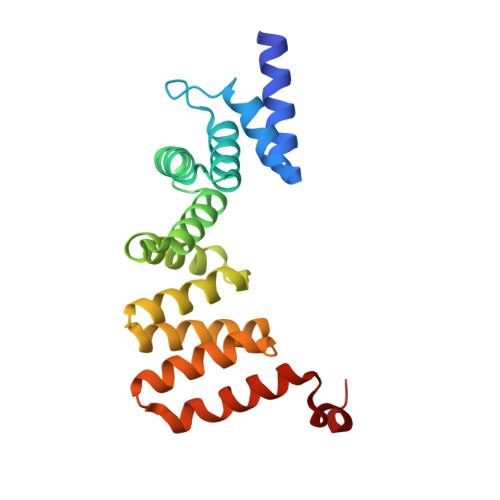
Self-recognition mechanism of MamA, a magnetosome-associated TPR-containing protein, promotes complex assembly
Zeytuni, N., Ozyamak, E., Ben-Harush, K., Davidov, G., Levin, M., Gat, Y., Moyal, T., Brik, A., Komeili, A., Zarivach, R.(2011) Proc Natl Acad Sci U S A 108: E480-E487
- PubMed: 21784982 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1103367108
- Primary Citation Related Structures:
3AS4, 3AS5, 3AS8, 3ASD, 3ASF, 3ASG, 3ASH - PubMed Abstract:
The magnetosome, a biomineralizing organelle within magnetotactic bacteria, allows their navigation along geomagnetic fields. Magnetosomes are membrane-bound compartments containing magnetic nanoparticles and organized into a chain within the cell, the assembly and biomineralization of magnetosomes are controlled by magnetosome-associated proteins. Here, we describe the crystal structures of the magnetosome-associated protein, MamA, from Magnetospirillum magneticum AMB-1 and Magnetospirillum gryphiswaldense MSR-1. MamA folds as a sequential tetra-trico-peptide repeat (TPR) protein with a unique hook-like shape. Analysis of the MamA structures indicates two distinct domains that can undergo conformational changes. Furthermore, structural analysis of seven crystal forms verified that the core of MamA is not affected by crystallization conditions and identified three protein-protein interaction sites, namely a concave site, a convex site, and a putative TPR repeat. Additionally, relying on transmission electron microscopy and size exclusion chromatography, we show that highly stable complexes form upon MamA homooligomerization. Disruption of the MamA putative TPR motif or N-terminal domain led to protein mislocalization in vivo and prevented MamA oligomerization in vitro. We, therefore, propose that MamA self-assembles through its putative TPR motif and its concave site to create a large homooligomeric scaffold which can interact with other magnetosome-associated proteins via the MamA convex site. We discuss the structural basis for TPR homooligomerization that allows the proper function of a prokaryotic organelle.
- Department of Life Sciences, Ben Gurion University of the Negev, P.O.B. 653, Beer-Sheva 84105, Israel.
Organizational Affiliation: