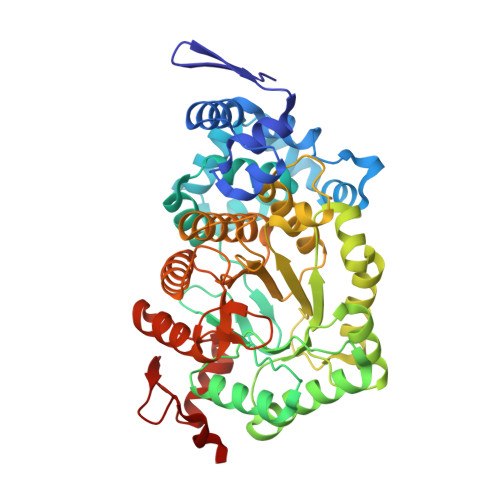
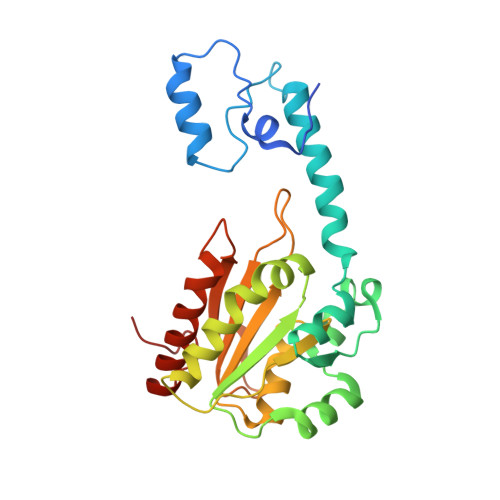
How coenzyme B12-dependent ethanolamine ammonia-lyase deals with both enantiomers of 2-amino-1-propanol as substrates: structure-based rationalization.
Shibata, N., Higuchi, Y., Toraya, T.(2011) Biochemistry 50: 591-598
- PubMed: 21142024 Search on PubMed
- DOI: https://doi.org/10.1021/bi101696h
- Primary Citation Related Structures:
3ANY, 3AO0 - PubMed Abstract:
Coenzyme B(12)-dependent ethanolamine ammonia-lyase acts on both enantiomers of the substrate 2-amino-1-propanol [Diziol, P., et al. (1980) Eur. J. Biochem. 106, 211-224]. To rationalize this apparent lack of stereospecificity and the enantiomer-specific stereochemical courses of the deamination, we analyzed the X-ray structures of enantiomer-bound forms of the enzyme-cyanocobalamin complex. The lower affinity for the (R)-enantiomer may be due to the conformational change of the Valα326 side chain of the enzyme. In a manner consistent with the reported experimental results, we can predict that the pro-S hydrogen atom on C1 is abstracted by the adenosyl radical from both enantiomeric substrates, because it is the nearest one in both enantiomer-bound forms. We also predicted that the NH(2) group migrates from C2 to C1 by a suprafacial shift, with inversion of configuration at C1 for both enantiomeric substrates, although the absolute configuration of the 1-amino-1-propanol intermediate is not yet known. Reported labeling experiments demonstrate that (R)-2-amino-1-propanol is deaminated by the enzyme with inversion of configuration at C2, whereas the (S)-enantiomer is deaminated with retention. By taking these results into consideration, we can predict the rotameric radical intermediate from the (S)-enantiomer undergoes flipping to the rotamer from the (R)-enantiomer before the hydrogen back-abstraction. This suggests the preference of the enzyme active site for the rotamer from the (R)-enantiomer in equilibration. This preference might be explained in terms of the steric repulsion of the (S)-enantiomer-derived product radical at C3 with the Pheα329 and Leuα402 residues.
- Department of Life Science, Graduate School of Life Science, University of Hyogo, 3-2-1 Koto, Kamigori-cho, Ako-gun, Hyogo, Japan. shibach@sci.u-hyogo.ac.jp
Organizational Affiliation: