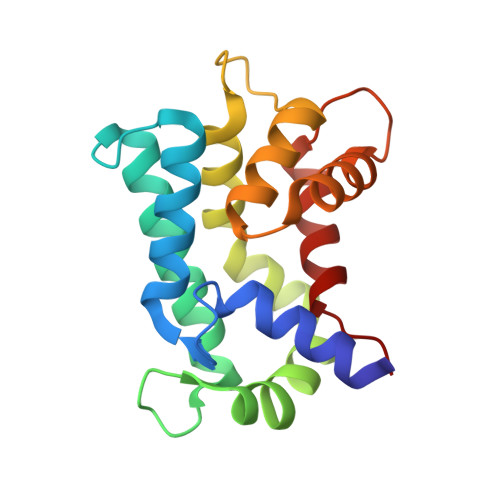
Structural basis for prokaryotic calciummediated regulation by a Streptomyces coelicolor calcium binding protein
Zhao, X., Pang, H., Wang, S., Zhou, W., Yang, K., Bartlam, M.(2010) Protein Cell 1: 771-779
- PubMed: 21203918 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s13238-010-0085-z
- Primary Citation Related Structures:
3AKA, 3AKB - PubMed Abstract:
The important and diverse regulatory roles of Ca(2+) in eukaryotes are conveyed by the EF-hand containing calmodulin superfamily. However, the calcium-regulatory proteins in prokaryotes are still poorly understood. In this study, we report the three-dimensional structure of the calcium-binding protein from Streptomyces coelicolor, named CabD, which shares low sequence homology with other known helix-loop-helix EF-hand proteins. The CabD structure should provide insights into the biological role of the prokaryotic calcium-binding proteins. The unusual structural features of CabD compared with prokaryotic EF-hand proteins and eukaryotic sarcoplasmic calcium-binding proteins, including the bending conformation of the first C-terminal α-helix, unpaired ligand-binding EF-hands and the lack of the extreme C-terminal loop region, suggest it may have a distinct and significant function in calcium-mediated bacterial physiological processes, and provide a structural basis for potential calcium-mediated regulatory roles in prokaryotes.
- Laboratory of Structural Biology, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: