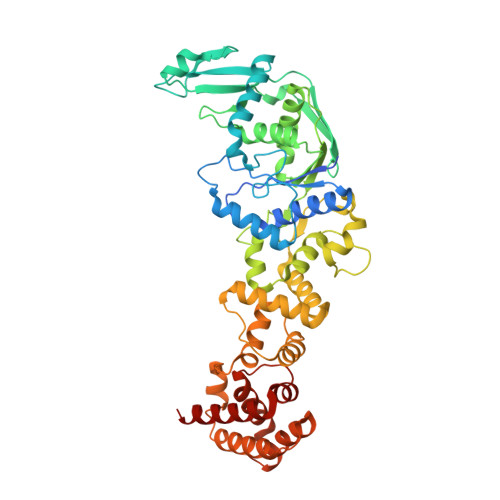
Structure of nondiscriminating glutamyl-tRNA synthetase from Thermotoga maritima
Ito, T., Kiyasu, N., Matsunaga, R., Takahashi, S., Yokoyama, S.(2010) Acta Crystallogr D Biol Crystallogr 66: 813-820
- PubMed: 20606262 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444910019086
- Primary Citation Related Structures:
3AFH - PubMed Abstract:
Aminoacyl-tRNA synthetases produce aminoacyl-tRNAs from the substrate tRNA and its cognate amino acid with the aid of ATP. Two types of glutamyl-tRNA synthetase (GluRS) have been discovered: discriminating GluRS (D-GluRS) and nondiscriminating GluRS (ND-GluRS). D-GluRS glutamylates tRNA(Glu) only, while ND-GluRS glutamylates both tRNA(Glu) and tRNA(Gln). ND-GluRS produces the intermediate Glu-tRNA(Gln), which is converted to Gln-tRNA(Gln) by Glu-tRNA(Gln) amidotransferase. Two GluRS homologues from Thermotoga maritima, TM1875 and TM1351, have been biochemically characterized and it has been clarified that only TM1875 functions as an ND-GluRS. Furthermore, the crystal structure of the T. maritima ND-GluRS, TM1875, was determined in complex with a Glu-AMP analogue at 2.0 A resolution. The T. maritima ND-GluRS contains a characteristic structure in the connective-peptide domain, which is inserted into the catalytic Rossmann-fold domain. The glutamylation ability of tRNA(Gln) by ND-GluRS was measured in the presence of the bacterial Glu-tRNA(Gln) amidotransferase GatCAB. Interestingly, the glutamylation efficiency was not affected even in the presence of excess GatCAB. Therefore, GluRS avoids competition with GatCAB and glutamylates tRNA(Gln).
- Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, Bunkyo-ku, Tokyo, Japan.
Organizational Affiliation: