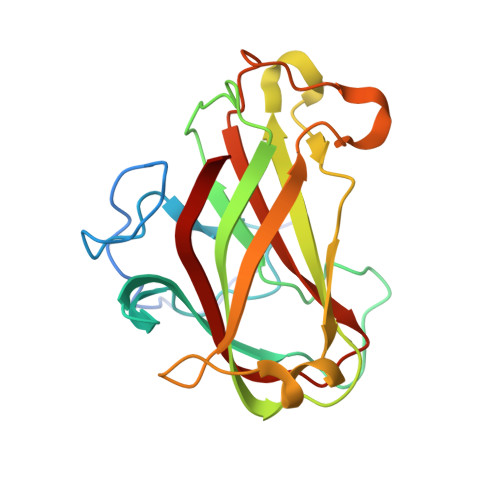
Recognition of cellooligosaccharides by a family 28 carbohydrate-binding module.
Tsukimoto, K., Takada, R., Araki, Y., Suzuki, K., Karita, S., Wakagi, T., Shoun, H., Watanabe, T., Fushinobu, S.(2010) FEBS Lett 584: 1205-1211
- PubMed: 20159017 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2010.02.027
- Primary Citation Related Structures:
3ACF, 3ACG, 3ACH, 3ACI - PubMed Abstract:
The crystal structures of a carbohydrate-binding module (CBM) family 28 domain of endoglucanase Cel5A from Clostridium josui have been determined in ligand-free and complex forms with cellobiose, cellotetraose, and cellopentaose as the first complex structures of this family. In the cleft of a beta-sandwich fold, the ligands are recognized by stacking interactions and hydrogen bonds. Conformations of the bound cellooligosaccharides are similar to those in crystals and solution but clearly different from the cellulose structure. Interestingly, the glucan chain bound on CBM28 is in the opposite direction of that bound to CBM17, although these families share significant structural similarity.
- Department of Biotechnology, The University of Tokyo, Tokyo, Japan.
Organizational Affiliation: