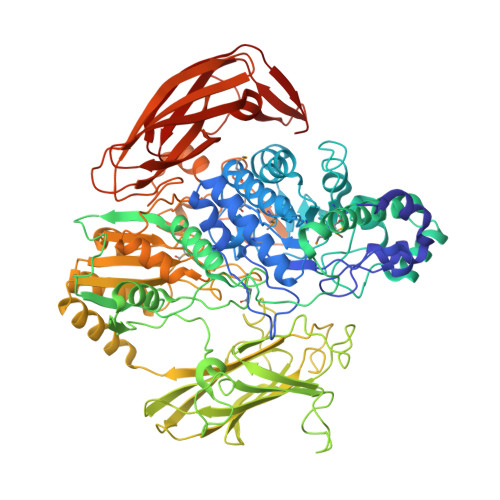
Role of a PA14 domain in determining substrate specificity of a glycoside hydrolase family 3 beta-glucosidase from Kluyveromyces marxianus.
Yoshida, E., Hidaka, M., Fushinobu, S., Koyanagi, T., Minami, H., Tamaki, H., Kitaoka, M., Katayama, T., Kumagai, H.(2010) Biochem J 431: 39-49
- PubMed: 20662765
- DOI: https://doi.org/10.1042/BJ20100351
- Primary Citation Related Structures:
3ABZ, 3AC0 - PubMed Abstract:
β-Glucosidase from Kluyveromyces marxianus (KmBglI) belongs to the GH3 (glycoside hydrolase family 3). The enzyme is particularly unusual in that a PA14 domain (pf07691), for which a carbohydrate-binding role has been claimed, is inserted into the catalytic core sequence. In the present study, we determined the enzymatic properties and crystal structure of KmBglI in complex with glucose at a 2.55 A (1 A=0.1 nm) resolution. A striking characteristic of KmBglI was that the enzyme activity is essentially limited to disaccharides, and when trisaccharides were used as the substrates the activity was drastically decreased. This chain-length specificity is in sharp contrast with the preferred action on oligosaccharides of barley β-D-glucan glucohydrolase (ExoI), which does not have a PA14 domain insertion. The structure of subsite (-1) of KmBglI is almost identical with that of Thermotoga neapolitana β-glucosidase and is also similar to that of ExoI, however, the structures of subsite (+1) significantly differ among them. In KmBglI, the loops extending from the PA14 domain cover the catalytic pocket to form subsite (+1), and hence simultaneously become a steric hindrance that could limit the chain length of the substrates to be accommodated. Mutational studies demonstrated the critical role of the loop regions in determining the substrate specificity. The active-site formation mediated by the PA14 domain of KmBglI invokes α-complementation of β-galactosidase exerted by its N-terminal domain, to which the PA14 domain shows structural resemblance. The present study is the first which reveals the structural basis of the interaction between the PA14 domain and a carbohydrate.
- Research Institute for Bioresources and Biotechnology, Ishikawa Prefectural University, Nonoichi, Ishikawa 921-8836, Japan.
Organizational Affiliation: