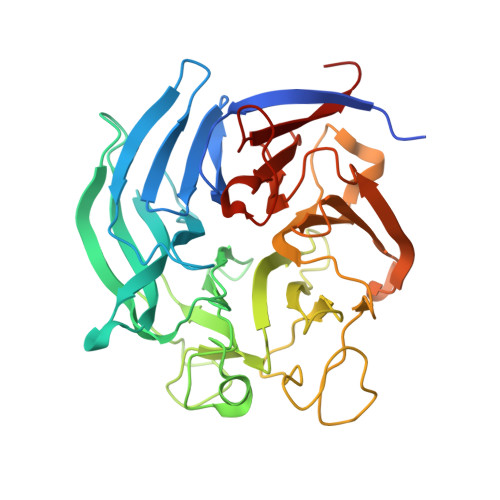
Catalytic properties and crystal structure of quinoprotein aldose sugar dehydrogenase from hyperthermophilic archaeon Pyrobaculum aerophilum
Sakuraba, H., Yokono, K., Yoneda, K., Watanabe, A., Asada, Y., Satomura, T., Yabutani, T., Motonaka, J., Ohshima, T.(2010) Arch Biochem Biophys 502: 81-88
- PubMed: 20692227
- DOI: https://doi.org/10.1016/j.abb.2010.08.002
- Primary Citation Related Structures:
3A9G, 3A9H - PubMed Abstract:
We identified a gene encoding a soluble quinoprotein glucose dehydrogenase homologue in the hyperthermophilic archaeon Pyrobaculum aerophilum. The gene was overexpressed in Escherichia coli, after which its product was purified and characterized. The enzyme was extremely thermostable, and the activity of the pyrroloquinoline quinone (PQQ)-bound holoenzyme was not lost after incubation at 100 degrees C for 10 min. The crystal structure of the enzyme was determined in both the apoform and as the PQQ-bound holoenzyme. The overall fold of the P. aerophilum enzyme showed significant similarity to that of soluble quinoprotein aldose sugar dehydrogenase (Asd) from E. coli. However, clear topological differences were observed in the two long loops around the PQQ-binding sites of the two enzymes. Structural comparison revealed that the hyperthermostability of the P. aerophilum enzyme is likely attributable to the presence of an extensive aromatic pair network located around a beta-sheet involving N- and C-terminal beta-strands.
- Department of Applied Biological Science, Kagawa University, Miki-cho, Kita-gun, Japan.
Organizational Affiliation: