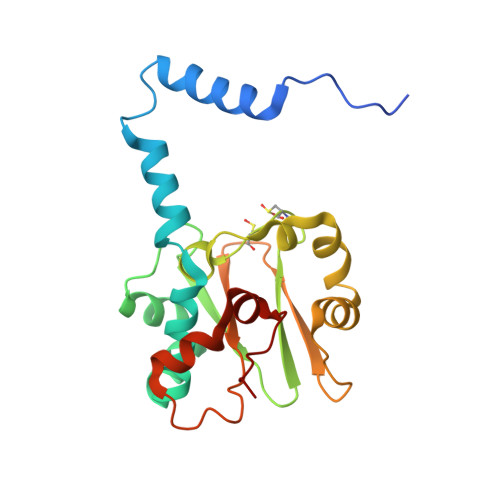
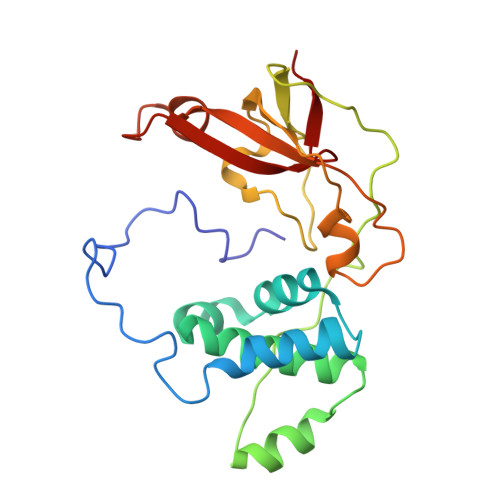
Kinetic and structural studies on roles of the serine ligand and a strictly conserved tyrosine residue in nitrile hydratase
Yamanaka, Y., Hashimoto, K., Ohtaki, A., Noguchi, K., Yohda, M., Odaka, M.(2010) J Biol Inorg Chem 15: 655-665
- PubMed: 20221653 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-010-0632-3
- Primary Citation Related Structures:
3A8G, 3A8H, 3A8L, 3A8M, 3A8O - PubMed Abstract:
Nitrile hydratases (NHase), which catalyze the hydration of nitriles to amides, have an unusual Fe(3+) or Co(3+) center with two modified Cys ligands: cysteine sulfininate (Cys-SO(2) (-)) and either cysteine sulfenic acid or cysteine sulfenate [Cys-SO(H)]. Two catalytic mechanisms have been proposed. One is that the sulfenyl oxygen activates a water molecule, enabling nucleophilic attack on the nitrile carbon. The other is that the Ser ligand ionizes the strictly conserved Tyr, activating a water molecule. Here, we characterized mutants of Fe-type NHase from Rhodococcus erythropolis N771, replacing the Ser and Tyr residues, alphaS113A and betaY72F. The alphaS113A mutation partially affected catalytic activity and did not change the pH profiles of the kinetic parameters. UV-vis absorption spectra indicated that the electronic state of the Fe center was altered by the alphaS113A mutation, but the changes could be prevented by a competitive inhibitor, n-butyric acid. The overall structure of the alphaS113A mutant was similar to that of the wild type, but significant changes were observed around the catalytic cavity. Like the UV-vis spectra, the changes were compensated by the substrate or product. The Ser ligand is important for the structure around the catalytic cavity, but is not essential for catalysis. The betaY72F mutant exhibited no activity. The structure of the betaY72F mutant was highly conserved but was found to be the inactivated state, with alphaCys114-SO(H) oxidized to Cys-SO(2) (-), suggesting that betaTyr72 affected the electronic state of the Fe center. The catalytic mechanism is discussed on the basis of the results obtained.
- Department of Biotechnology and Life Science, Tokyo University of Agriculture and Technology, Naka-cho, Koganei, Japan.
Organizational Affiliation: