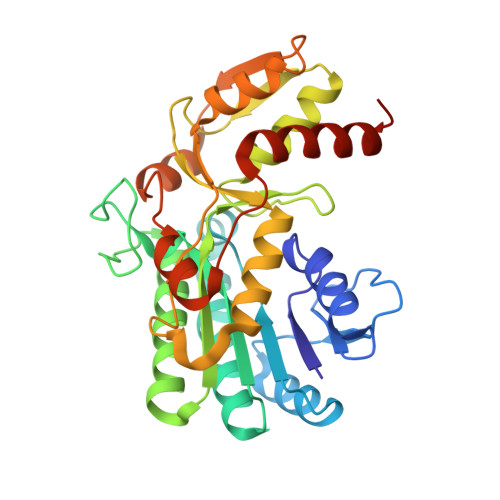
Crystal Structure of Binary and Ternary Complexes of Archaeal UDP-galactose 4-Epimerase-like L-Threonine Dehydrogenase from Thermoplasma volcanium.
Yoneda, K., Sakuraba, H., Araki, T., Ohshima, T.(2012) J Biological Chem 287: 12966-12974
- PubMed: 22374996 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.336958
- Primary Citation Related Structures:
3A1N, 3A4V, 3A9W, 3AJR - PubMed Abstract:
A gene from the thermophilic archaeon Thermoplasma volcanium encoding an L-threonine dehydrogenase (L-ThrDH) with a predicted amino acid sequence that was remarkably similar to the sequence of UDP-galactose 4-epimerase (GalE) was overexpressed in Escherichia coli, and its product was purified and characterized. The expressed enzyme was moderately thermostable, retaining more than 90% of its activity after incubation for 10 min at up to 70 °C. The catalytic residue was assessed using site-directed mutagenesis, and Tyr(137) was found to be essential for catalysis. To clarify the structural basis of the catalytic mechanism, four different crystal structures were determined using the molecular replacement method: L-ThrDH-NAD(+), L-ThrDH in complex with NAD(+) and pyruvate, Y137F mutant in complex with NAD(+) and L-threonine, and Y137F in complex with NAD(+) and L-3-hydroxynorvaline. Each monomer consisted of a Rossmann-fold domain and a C-terminal catalytic domain, and the fold of the catalytic domain showed notable similarity to that of the GalE-like L-ThrDH from the psychrophilic bacterium Flavobacterium frigidimaris KUC-1. The substrate binding model suggests that the reaction proceeds through abstraction of the β-hydroxyl hydrogen of L-threonine via direct proton transfer driven by Tyr(137). The factors contributing to the thermostability of T. volcanium L-ThrDH were analyzed by comparing its structure to that of F. frigidimaris L-ThrDH. This comparison showed that the presence of extensive inter- and intrasubunit ion pair networks are likely responsible for the thermostability of T. volcanium L-ThrDH. This is the first description of the molecular basis for the substrate recognition and thermostability of a GalE-like L-ThrDH.
- Department of Bioscience, School of Agriculture, Tokai University, Aso, Kumamoto, 869-1404, Japan.
Organizational Affiliation: