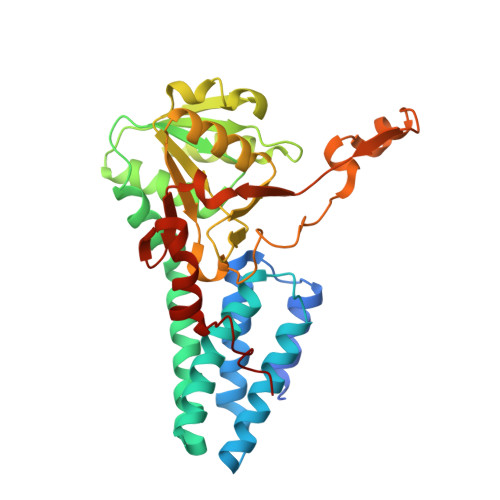
Dynamic, ligand-dependent conformational change triggers reaction of ribose-1,5-bisphosphate isomerase from Thermococcus kodakarensis KOD1
Nakamura, A., Fujihashi, M., Aono, R., Sato, T., Nishiba, Y., Yoshida, S., Yano, A., Atomi, H., Imanaka, T., Miki, K.(2012) J Biological Chem 287: 20784-20796
- PubMed: 22511789 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.349423
- Primary Citation Related Structures:
3A11, 3A9C, 3VM6 - PubMed Abstract:
Ribose-1,5-bisphosphate isomerase (R15Pi) is a novel enzyme recently identified as a member of an AMP metabolic pathway in archaea. The enzyme converts d-ribose 1,5-bisphosphate into ribulose 1,5-bisphosphate, providing the substrate for archaeal ribulose-1,5-bisphosphate carboxylase/oxygenases. We here report the crystal structures of R15Pi from Thermococcus kodakarensis KOD1 (Tk-R15Pi) with and without its substrate or product. Tk-R15Pi is a hexameric enzyme formed by the trimerization of dimer units. Biochemical analyses show that Tk-R15Pi only accepts the α-anomer of d-ribose 1,5-bisphosphate and that Cys(133) and Asp(202) residues are essential for ribulose 1,5-bisphosphate production. Comparison of the determined structures reveals that the unliganded and product-binding structures are in an open form, whereas the substrate-binding structure adopts a closed form, indicating domain movement upon substrate binding. The conformational change to the closed form optimizes active site configuration and also isolates the active site from the solvent, which may allow deprotonation of Cys(133) and protonation of Asp(202) to occur. The structural features of the substrate-binding form and biochemical evidence lead us to propose that the isomerase reaction proceeds via a cis-phosphoenolate intermediate.
- Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto 606-8502, Japan.
Organizational Affiliation: