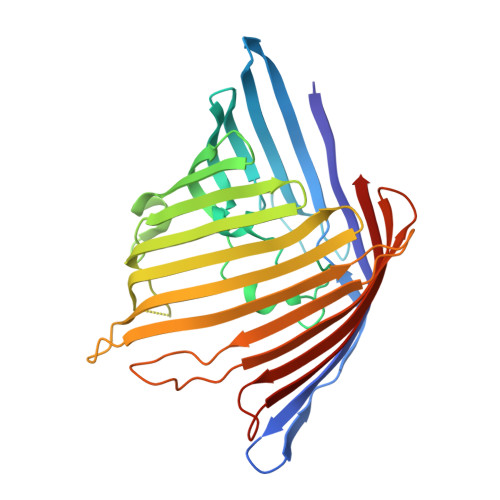
Crystallographic analysis of Neisseria meningitidis PorB extracellular loops potentially implicated in TLR2 recognition.
Kattner, C., Toussi, D.N., Zaucha, J., Wetzler, L.M., Ruppel, N., Zachariae, U., Massari, P., Tanabe, M.(2014) J Struct Biol 185: 440-447
- PubMed: 24361688 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jsb.2013.12.006
- Primary Citation Related Structures:
3WI4, 3WI5 - PubMed Abstract:
Among all Neisseriae species, Neisseria meningitidis and Neisseria gonorrhoeae are the only human pathogens, causative agents of bacterial meningitis and gonorrhoea, respectively. PorB, a pan-Neisseriae trimeric porin that mediates diffusive transport of essential molecules across the bacterial outer membrane, is also known to activate host innate immunity via Toll-like receptor 2 (TLR2)-mediated signaling. The molecular mechanism of PorB binding to TLR2 is not known, but it has been hypothesized that electrostatic interactions contribute to ligand/receptor binding. Strain-specific sequence variability in the surface-exposed loops of PorB which are potentially implicated in TLR2 binding, may explain the difference in TLR2-mediated cell activation in vitro by PorB homologs from the commensal Neisseriae lactamica and the pathogen N. meningitidis. Here, we report a comparative structural analysis of PorB from N. meningitidis serogroup B strain 8765 (63% sequence homology with PorB from N. meningitidis serogroup W135) and a mutant in which amino acid substitutions in the extracellular loop 7 lead to significantly reduced TLR2-dependent activity in vitro. We observe that this mutation both alters the loop conformation and causes dramatic changes of electrostatic surface charge, both of which may affect TLR2 recognition and signaling.
- HALOmem, Membrane Protein Biochemistry, Martin-Luther-University Halle-Wittenberg, Halle (Saale), Germany.
Organizational Affiliation: