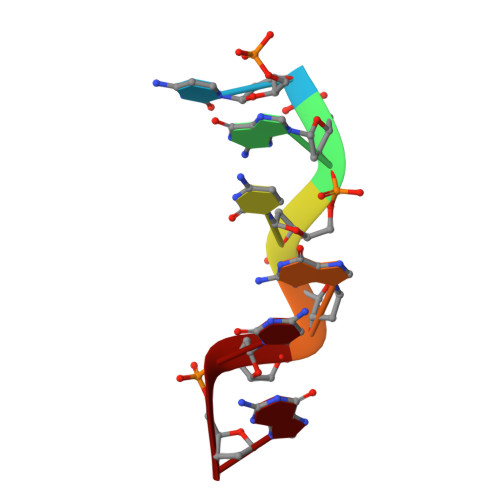
Structural variability and new intermolecular interactions of Z-DNA in crystals of d(pCpGpCpGpCpG).
Malinina, L., Tereshko, V., Ivanova, E., Subirana, J.A., Zarytova, V., Nekrasov, Y.(1998) Biophys J 74: 2482-2490
- PubMed: 9591674 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/S0006-3495(98)77956-1
- Primary Citation Related Structures:
390D, 391D, 392D - PubMed Abstract:
We have determined the single crystal x-ray structure of the synthetic DNA hexamer d(pCpGpCpGpCpG) in two different crystal forms. The hexamer pCGCGCG has the Z-DNA conformation and in both cases the asymmetric unit contains more than one Z-DNA duplex. Crystals belong to the space group C222(1) with a = 69.73, b = 52.63, and c = 26.21 A, and to the space group P2(1) with a = 49.87, b = 41.26, c = 21.91 A, and gamma = 97.12 degrees. Both crystals show new crystal packing modes. The molecules also show striking new features when compared with previously determined Z-DNA structures: 1) the bases in one duplex have a large inclination with respect to the helical axis, which alters the overall shape of the molecule. 2) Some cytosine nitrogens interact by hydrogen bonding with phosphates in neighbor molecules. Similar base-phosphate interactions had been previously detected in some B-DNA crystals. 3) Basepair stacking between the ends of neighbor molecules is variable and no helical continuity is maintained between contiguous hexamer duplexes.
- Engelhardt Institute of Molecular Biology RAS, Moscow, Russia. lucy@crystal.eimb.rssi.ru
Organizational Affiliation: