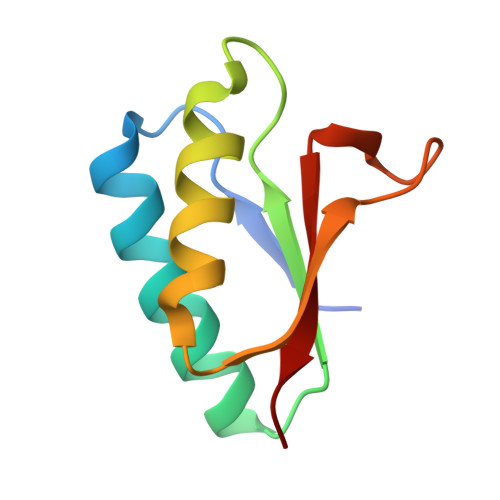
Crystal structure of MutS2 endonuclease domain and the mechanism of homologous recombination suppression
Fukui, K., Nakagawa, N., Kitamura, Y., Nishida, Y., Masui, R., Kuramitsu, S.(2008) J Biological Chem 283: 33417-33427
- PubMed: 18838375 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M806755200
- Primary Citation Related Structures:
2ZQE - PubMed Abstract:
DNA recombination events need to be strictly regulated, because an increase in the recombinational frequency causes unfavorable alteration of genetic information. Recent studies revealed the existence of a novel anti-recombination enzyme, MutS2. However, the mechanism by which MutS2 inhibits homologous recombination has been unknown. Previously, we found that Thermus thermophilus MutS2 (ttMutS2) harbors an endonuclease activity and that this activity is confined to the C-terminal domain, whose amino acid sequence is widely conserved in a variety of proteins with unknown function from almost all organisms ranging from bacteria to man. In this study, we determined the crystal structure of the ttMutS2 endonuclease domain at 1.7-angstroms resolution, which resembles the structure of the DNase I-like catalytic domain of Escherichia coli RNase E, a sequence-nonspecific endonuclease. The N-terminal domain of ttMutS2, however, recognized branched DNA structures, including the Holliday junction and D-loop structure, a primary intermediate in homologous recombination. The full-length of ttMutS2 digested the branched DNA structures at the junction. These results indicate that ttMutS2 suppresses homologous recombination through a novel mechanism involving resolution of early intermediates.
- RIKEN SPring-8 Center, Harima Institute, 1-1-1 Kouto, Sayo-gun, Hyogo, 679-5148, Japan.
Organizational Affiliation: