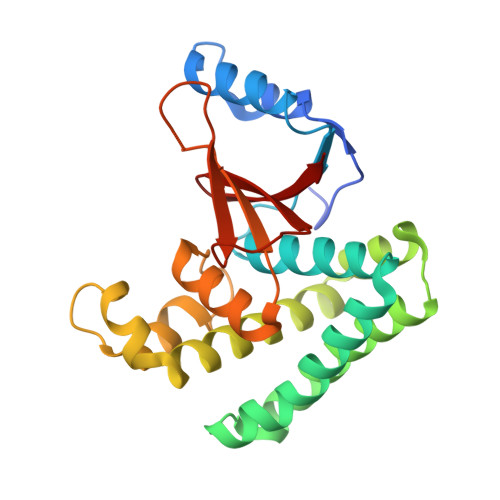
Structural basis and specificity of human otubain 1-mediated deubiquitination.
Edelmann, M.J., Iphofer, A., Akutsu, M., Altun, M., di Gleria, K., Kramer, H.B., Fiebiger, E., Dhe-Paganon, S., Kessler, B.M.(2009) Biochem J 418: 379-390
- PubMed: 18954305 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20081318
- Primary Citation Related Structures:
2ZFY - PubMed Abstract:
OTUB (otubain) 1 is a human deubiquitinating enzyme that is implicated in mediating lymphocyte antigen responsiveness, but whose molecular function is generally not well defined. A structural analysis of OTUB1 shows differences in accessibility to the active site and in surface properties of the substrate-binding regions when compared with its close homologue, OTUB2, suggesting variations in regulatory mechanisms and substrate specificity. Biochemical analysis reveals that OTUB1 has a preference for cleaving Lys(48)-linked polyubiquitin chains over Lys(63)-linked polyubiquitin chains, and it is capable of cleaving NEDD8 (neural-precursor-cell-expressed developmentally down-regulated 8), but not SUMO (small ubiquitin-related modifier) 1/2/3 and ISG15 (interferon-stimulated gene 15) conjugates. A functional comparison of OTUB1 and OTUB2 indicated a differential reactivity towards ubiquitin-based active-site probes carrying a vinyl methyl ester, a 2-chloroethyl or a 2-bromoethyl group at the C-terminus. Mutational analysis suggested that a narrow P1' site, as observed in OTUB1, correlates with its ability to preferentially cleave Lys(48)-linked ubiquitin chains. Analysis of cellular interaction partners of OTUB1 by co-immunoprecipitation and MS/MS (tandem mass spectrometry) experiments demonstrated that FUS [fusion involved in t(12;6) in malignant liposarcoma; also known as TLS (translocation in liposarcoma) or CHOP (CCAAT/enhancer-binding protein homologous protein)] and RACK1 [receptor for activated kinase 1; also known as GNB2L1 (guanine-nucleotide-binding protein beta polypeptide 2-like 1)] are part of OTUB1-containing complexes, pointing towards a molecular function of this deubiquitinating enzyme in RNA processing and cell adhesion/morphology.
- Henry Wellcome Building for Molecular Physiology, Department of Clinical Medicine, University of Oxford, Roosevelt Drive, Oxford OX37BN, UK.
Organizational Affiliation: